Toward a Self-Updating Platform for Estimating Rates of Speciation and Migration, Ages, and Relationships of Taxa
- PMID: 27616324
- PMCID: PMC5410925
- DOI: 10.1093/sysbio/syw066
Toward a Self-Updating Platform for Estimating Rates of Speciation and Migration, Ages, and Relationships of Taxa
Abstract
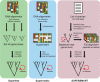
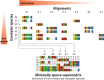
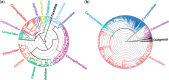
Rapidly growing biological data-including molecular sequences and fossils-hold an unprecedented potential to reveal how evolutionary processes generate and maintain biodiversity. However, researchers often have to develop their own idiosyncratic workflows to integrate and analyze these data for reconstructing time-calibrated phylogenies. In addition, divergence times estimated under different methods and assumptions, and based on data of various quality and reliability, should not be combined without proper correction. Here we introduce a modular framework termed SUPERSMART (Self-Updating Platform for Estimating Rates of Speciation and Migration, Ages, and Relationships of Taxa), and provide a proof of concept for dealing with the moving targets of evolutionary and biogeographical research. This framework assembles comprehensive data sets of molecular and fossil data for any taxa and infers dated phylogenies using robust species tree methods, also allowing for the inclusion of genomic data produced through next-generation sequencing techniques. We exemplify the application of our method by presenting phylogenetic and dating analyses for the mammal order Primates and for the plant family Arecaceae (palms). We believe that this framework will provide a valuable tool for a wide range of hypothesis-driven research questions in systematics, biogeography, and evolution. SUPERSMART will also accelerate the inference of a "Dated Tree of Life" where all node ages are directly comparable. [Bayesian phylogenetics; data mining; divide-and-conquer methods; GenBank; multilocus multispecies coalescent; next-generation sequencing; palms; primates; tree calibration.].
© The Author(s) 2016. Published by Oxford University Press, on behalf of the Society of Systematic Biologists.
Figures



References
-
- Antonelli A. Forthcoming. Advancing biodiversity research: comparative biogeography, big data, and common myths. Sci. Danica.
-
- Bacon C.D., Michonneau F., Henderson A.J. McKenna M.J., Milroy A.M., Simmons M.P.. 2013.. Geographic and taxonomic disparities in species diversity: dispersal and diversification rates across wallace’s line. Evolution 67:2058–2071. - PubMed
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

