A Comparison of Deterministic and Stochastic Modeling Approaches for Biochemical Reaction Systems: On Fixed Points, Means, and Modes
- PMID: 27630669
- PMCID: PMC5005346
- DOI: 10.3389/fgene.2016.00157
A Comparison of Deterministic and Stochastic Modeling Approaches for Biochemical Reaction Systems: On Fixed Points, Means, and Modes
Abstract
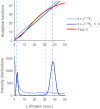
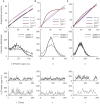
In the mathematical modeling of biochemical reactions, a convenient standard approach is to use ordinary differential equations (ODEs) that follow the law of mass action. However, this deterministic ansatz is based on simplifications; in particular, it neglects noise, which is inherent to biological processes. In contrast, the stochasticity of reactions is captured in detail by the discrete chemical master equation (CME). Therefore, the CME is frequently applied to mesoscopic systems, where copy numbers of involved components are small and random fluctuations are thus significant. Here, we compare those two common modeling approaches, aiming at identifying parallels and discrepancies between deterministic variables and possible stochastic counterparts like the mean or modes of the state space probability distribution. To that end, a mathematically flexible reaction scheme of autoregulatory gene expression is translated into the corresponding ODE and CME formulations. We show that in the thermodynamic limit, deterministic stable fixed points usually correspond well to the modes in the stationary probability distribution. However, this connection might be disrupted in small systems. The discrepancies are characterized and systematically traced back to the magnitude of the stoichiometric coefficients and to the presence of nonlinear reactions. These factors are found to synergistically promote large and highly asymmetric fluctuations. As a consequence, bistable but unimodal, and monostable but bimodal systems can emerge. This clearly challenges the role of ODE modeling in the description of cellular signaling and regulation, where some of the involved components usually occur in low copy numbers. Nevertheless, systems whose bimodality originates from deterministic bistability are found to sustain a more robust separation of the two states compared to bimodal, but monostable systems. In regulatory circuits that require precise coordination, ODE modeling is thus still expected to provide relevant indications on the underlying dynamics.
Keywords: bimodality; bistability; chemical master equation; gene expression; ordinary differential equations; protein bursts.
Figures

References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

