Imaging flow cytometry analysis of intracellular pathogens
- PMID: 27642004
- PMCID: PMC5857943
- DOI: 10.1016/j.ymeth.2016.09.007
Imaging flow cytometry analysis of intracellular pathogens
Abstract
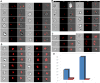
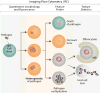
Imaging flow cytometry has been applied to address questions in infection biology, in particular, infections induced by intracellular pathogens. This methodology, which utilizes specialized analytic software makes it possible to analyze hundreds of quantified features for hundreds of thousands of individual cellular or subcellular events in a single experiment. Imaging flow cytometry analysis of host cell-pathogen interaction can thus quantitatively addresses a variety of biological questions related to intracellular infection, including cell counting, internalization score, and subcellular patterns of co-localization. Here, we provide an overview of recent achievements in the use of fluorescently labeled prokaryotic or eukaryotic pathogens in human cellular infections in analysis of host-pathogen interactions. Specifically, we give examples of Imagestream-based analysis of cell lines infected with Toxoplasma gondii or Mycobacterium tuberculosis. Furthermore, we illustrate the capabilities of imaging flow cytometry using a combination of standard IDEAS™ software and the more recently developed Feature Finder algorithm, which is capable of identifying statistically significant differences between researcher-defined image galleries. We argue that the combination of imaging flow cytometry with these software platforms provides a powerful new approach to understanding host control of intracellular pathogens.
Keywords: Cellular heterogeneity; Colocalization; Feature Finder; Fluorescent protein; Imaging flow cytometry; Intracellular pathogen; Mycobacteria tuberculosis; Phagosome maturation; Rab5; Rab7; Toxoplasma gondii.
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures




References
-
- Fuchs TM, Eisenreich W, Heesemann J, Goebel W. Metabolic adaptation of human pathogenic and related nonpathogenic bacteria to extra- and intracellular habitats. FEMS Microbiol. Rev. 2012;36:435–462. - PubMed
-
- Muskavitch MA, Barteneva N, Gubbels MJ. Chemogenomics and parasitology: small molecules and cell-based assays to study infectious processes. Comb. Chem. High Throughput Screen. 2008;11:624–646. - PubMed
-
- Toma C, Okura N, Takayama C, Suzuki T. Characteristic features of intracellular pathogenic Leptospira in infected murine macrophages. Cell. Microbiol. 2011;13:1783–1792. - PubMed
-
- Bailo N, Cosson P, Charette SJ, Paquet VE, Doublet P, Letourneur F. Defective lysosome maturation and Legionella pneumophila replication in Dictostelium cells mutant for the Arf GAP ACAP-A. J. Cell Sci. 2014;127:4702–4713. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

