Mass spectrometry as a quantitative tool in plant metabolomics
- PMID: 27644967
- PMCID: PMC5031636
- DOI: 10.1098/rsta.2015.0370
Mass spectrometry as a quantitative tool in plant metabolomics
Abstract
Metabolomics is a research field used to acquire comprehensive information on the composition of a metabolite pool to provide a functional screen of the cellular state. Studies of the plant metabolome include the analysis of a wide range of chemical species with very diverse physico-chemical properties, and therefore powerful analytical tools are required for the separation, characterization and quantification of this vast compound diversity present in plant matrices. In this review, challenges in the use of mass spectrometry (MS) as a quantitative tool in plant metabolomics experiments are discussed, and important criteria for the development and validation of MS-based analytical methods provided.This article is part of the themed issue 'Quantitative mass spectrometry'.
Keywords: analytical recoveries; mass spectrometry; matrix effects; metabolite quantification; method validation; plant metabolomics.
© 2016 The Author(s).
Figures


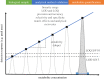
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
