Inference of gene regulation functions from dynamic transcriptome data
- PMID: 27652904
- PMCID: PMC5072840
- DOI: 10.7554/eLife.12188
Inference of gene regulation functions from dynamic transcriptome data
Abstract
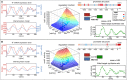
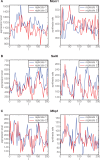
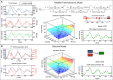
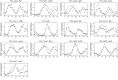
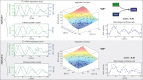
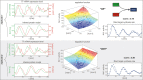
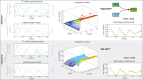
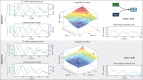
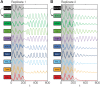
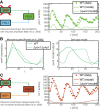
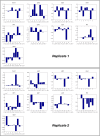
To quantify gene regulation, a function is required that relates transcription factor binding to DNA (input) to the rate of mRNA synthesis from a target gene (output). Such a 'gene regulation function' (GRF) generally cannot be measured because the experimental titration of inputs and simultaneous readout of outputs is difficult. Here we show that GRFs may instead be inferred from natural changes in cellular gene expression, as exemplified for the cell cycle in the yeast S. cerevisiae. We develop this inference approach based on a time series of mRNA synthesis rates from a synchronized population of cells observed over three cell cycles. We first estimate the functional form of how input transcription factors determine mRNA output and then derive GRFs for target genes in the CLB2 gene cluster that are expressed during G2/M phase. Systematic analysis of additional GRFs suggests a network architecture that rationalizes transcriptional cell cycle oscillations. We find that a transcription factor network alone can produce oscillations in mRNA expression, but that additional input from cyclin oscillations is required to arrive at the native behaviour of the cell cycle oscillator.
Keywords: S. cerevisiae; cell cycle; computational biology; gene regulation; network inference; quantitative biology; systems biology.
Conflict of interest statement
PC: Reviewing editor, eLife. The other authors declare that no competing interests exist.
Figures






Similar articles
-
Regulation of cell cycle-specific gene expression through cyclin-dependent kinase-mediated phosphorylation of the forkhead transcription factor Fkh2p.Mol Cell Biol. 2004 Nov;24(22):10036-46. doi: 10.1128/MCB.24.22.10036-10046.2004. Mol Cell Biol. 2004. PMID: 15509804 Free PMC article.
-
Transcriptional regulatory networks in Saccharomyces cerevisiae.Science. 2002 Oct 25;298(5594):799-804. doi: 10.1126/science.1075090. Science. 2002. PMID: 12399584
-
Two yeast forkhead genes regulate the cell cycle and pseudohyphal growth.Nature. 2000 Jul 6;406(6791):90-4. doi: 10.1038/35017581. Nature. 2000. PMID: 10894548
-
High functional overlap between MluI cell-cycle box binding factor and Swi4/6 cell-cycle box binding factor in the G1/S transcriptional program in Saccharomyces cerevisiae.Genetics. 2005 Sep;171(1):49-61. doi: 10.1534/genetics.105.044560. Epub 2005 Jun 18. Genetics. 2005. PMID: 15965243 Free PMC article.
-
The interplay between transcription and mRNA degradation in Saccharomyces cerevisiae.Microb Cell. 2017 Jul 3;4(7):212-228. doi: 10.15698/mic2017.07.580. Microb Cell. 2017. PMID: 28706937 Free PMC article. Review.
Cited by
-
Reconciling conflicting models for global control of cell-cycle transcription.Cell Cycle. 2017 Oct 18;16(20):1965-1978. doi: 10.1080/15384101.2017.1367073. Epub 2017 Sep 21. Cell Cycle. 2017. PMID: 28934013 Free PMC article.
-
Connecting virulence pathways to cell-cycle progression in the fungal pathogen Cryptococcus neoformans.Curr Genet. 2017 Oct;63(5):803-811. doi: 10.1007/s00294-017-0688-5. Epub 2017 Mar 6. Curr Genet. 2017. PMID: 28265742 Free PMC article. Review.
-
Reverse engineering highlights potential principles of large gene regulatory network design and learning.NPJ Syst Biol Appl. 2017 Jun 22;3:17. doi: 10.1038/s41540-017-0019-y. eCollection 2017. NPJ Syst Biol Appl. 2017. PMID: 28649444 Free PMC article.
-
Layers of regulation of cell-cycle gene expression in the budding yeast Saccharomyces cerevisiae.Mol Biol Cell. 2018 Nov 1;29(22):2644-2655. doi: 10.1091/mbc.E18-04-0255. Epub 2018 Sep 12. Mol Biol Cell. 2018. PMID: 30207828 Free PMC article.
-
The cell-cycle transcriptional network generates and transmits a pulse of transcription once each cell cycle.Cell Cycle. 2019 Feb;18(4):363-378. doi: 10.1080/15384101.2019.1570655. Epub 2019 Feb 5. Cell Cycle. 2019. PMID: 30668223 Free PMC article.
References
-
- Andrieu C, de Freitas N, Doucet A, Jordan MI. An introduction to MCMC for machine learning. Machine Learning. 2003;50:5–43. doi: 10.1023/A:1020281327116. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

