Neuronal Nuclear Membrane Budding Occurs during a Developmental Window Modulated by Torsin Paralogs
- PMID: 27653693
- PMCID: PMC5061049
- DOI: 10.1016/j.celrep.2016.08.044
Neuronal Nuclear Membrane Budding Occurs during a Developmental Window Modulated by Torsin Paralogs
Abstract
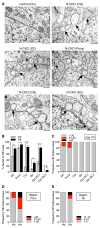
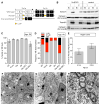
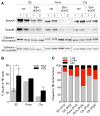
DYT1 dystonia is a neurodevelopmental disease that manifests during a discrete period of childhood. The disease is caused by impaired function of torsinA, a protein linked to nuclear membrane budding. The relationship of NE budding to neural development and CNS function is unclear, however, obscuring its potential role in dystonia pathogenesis. We find NE budding begins and resolves during a discrete neurodevelopmental window in torsinA null neurons in vivo. The developmental resolution of NE budding corresponds to increased torsinB protein, while ablating torsinB from torsinA null neurons prevents budding resolution and causes lethal neural dysfunction. Developmental changes in torsinB also correlate with NE bud formation in differentiating DYT1 embryonic stem cells, and overexpression of torsinA or torsinB rescues NE bud formation in this system. These findings identify a torsinA neurodevelopmental window that is essential for normal CNS function and have important implications for dystonia pathogenesis and therapeutics.
Copyright © 2016 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

