Dysregulated JAK2 expression by TrkC promotes metastasis potential, and EMT program of metastatic breast cancer
- PMID: 27654855
- PMCID: PMC5032000
- DOI: 10.1038/srep33899
Dysregulated JAK2 expression by TrkC promotes metastasis potential, and EMT program of metastatic breast cancer
Erratum in
-
Author Correction: Dysregulated JAK2 expression by TrkC promotes metastasis potential, and EMT program of metastatic breast cancer.Sci Rep. 2020 Jul 6;10(1):11103. doi: 10.1038/s41598-020-68128-6. Sci Rep. 2020. PMID: 32632170 Free PMC article.
Abstract
Metastatic breast cancers are aggressive tumors associated with high levels of epithelial-mesenchymal transition (EMT) markers, activation of IL6/JAK2/STAT3 and PI3K/AKT pathways for cell growth, mobility, invasion, metastasis, and CSC status. We identified a new molecular and functional network present in metastasis that regulates and coordinates with TrkC. Inhibition of SOCS3-mediated JAK2 degradation by TrkC increases total JAK2/STAT3 expression, and then leads to upregulation of Twist-1 through activation of JAK2/STAT3 cascade. Also, TrkC increases secretion and expression of IL-6, suggesting that this autocrine loop generated by TrkC maintains the mesenchymal state by continued activation of the JAK2/STAT3 cascade and upregulation of Twist expression. Moreover, TrkC interacts with the c-Src/Jak2 complex, which increases Twist-1 and Twist-2 levels via regulation of JAK2/STAT3 activation and JAK2/STAT3 expression. Furthermore, TrkC enhances metastatic potential of breast cancer via induction of EMT by upregulating Twist-1 and Twist-2. Additionally, TrkC significantly enhances the ability of breast cancer cells to form pulmonary metastases and primary tumor formation. Unexpectedly, we found that TrkC expression and clinical breast tumor pathological phenotypes show significant correlation. These findings suggest that TrkC plays a central role in tumorigenicity, metastasis, and self-renewal traits of metastatic breast cancer.
Figures



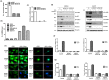



Similar articles
-
Induction of metastatic potential by TrkB via activation of IL6/JAK2/STAT3 and PI3K/AKT signaling in breast cancer.Oncotarget. 2015 Nov 24;6(37):40158-71. doi: 10.18632/oncotarget.5522. Oncotarget. 2015. PMID: 26515594 Free PMC article.
-
Cytoplasmic p27 promotes epithelial-mesenchymal transition and tumor metastasis via STAT3-mediated Twist1 upregulation.Oncogene. 2015 Oct;34(43):5447-59. doi: 10.1038/onc.2014.473. Epub 2015 Feb 16. Oncogene. 2015. PMID: 25684140 Free PMC article.
-
MEST induces Twist-1-mediated EMT through STAT3 activation in breast cancers.Cell Death Differ. 2019 Dec;26(12):2594-2606. doi: 10.1038/s41418-019-0322-9. Epub 2019 Mar 22. Cell Death Differ. 2019. PMID: 30903102 Free PMC article.
-
Roles of TrkC Signaling in the Regulation of Tumorigenicity and Metastasis of Cancer.Cancers (Basel). 2020 Jan 8;12(1):147. doi: 10.3390/cancers12010147. Cancers (Basel). 2020. PMID: 31936239 Free PMC article. Review.
-
Targeting Twist expression with small molecules.Medchemcomm. 2016 Dec 2;8(2):268-275. doi: 10.1039/c6md00561f. eCollection 2017 Feb 1. Medchemcomm. 2016. PMID: 30108743 Free PMC article. Review.
Cited by
-
Ribonucleotide reductase subunit M2 promotes proliferation and epithelial-mesenchymal transition via the JAK2/STAT3 signaling pathway in retinoblastoma.Bioengineered. 2021 Dec;12(2):12800-12811. doi: 10.1080/21655979.2021.2001241. Bioengineered. 2021. PMID: 34895038 Free PMC article.
-
TrkC promotes colorectal cancer growth and metastasis.Oncotarget. 2017 Jun 20;8(25):41319-41333. doi: 10.18632/oncotarget.17289. Oncotarget. 2017. PMID: 28455963 Free PMC article.
-
TrkC, a novel prognostic marker, induces and maintains cell survival and metastatic dissemination of Ewing sarcoma by inhibiting EWSR1-FLI1 degradation.Cell Death Dis. 2022 Sep 28;13(9):836. doi: 10.1038/s41419-022-05275-w. Cell Death Dis. 2022. PMID: 36171207 Free PMC article.
-
Therapeutic Potential of Ursolic Acid in Cancer and Diabetic Neuropathy Diseases.Int J Mol Sci. 2021 Nov 10;22(22):12162. doi: 10.3390/ijms222212162. Int J Mol Sci. 2021. PMID: 34830043 Free PMC article. Review.
-
TrkC-mediated inhibition of DJ-1 degradation is essential for direct regulation of pathogenesis of hepatocellular carcinoma.Cell Death Dis. 2022 Oct 6;13(10):850. doi: 10.1038/s41419-022-05298-3. Cell Death Dis. 2022. PMID: 36202793 Free PMC article.
References
-
- Hanahan D. & Weinberg R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011). - PubMed
-
- Batlle E. et al.. The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat Cell Biol 2, 84–89 (2000). - PubMed
-
- Cano A. et al.. The transcription factor snail controls epithelial-mesenchymal transitions by repressing E-cadherin expression. Nat Cell Biol 2, 76–83 (2000). - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

