To ∼P or Not to ∼P? Non-canonical activation by two-component response regulators
- PMID: 27656860
- PMCID: PMC5218973
- DOI: 10.1111/mmi.13532
To ∼P or Not to ∼P? Non-canonical activation by two-component response regulators
Abstract
Bacteria sense and respond to their environment through the use of two-component regulatory systems. The ability to adapt to a wide range of environmental stresses is directly related to the number of two-component systems an organism possesses. Recent advances in this area have identified numerous variations on the archetype systems that employ a sensor kinase and a response regulator. It is now evident that many orphan regulators that lack cognate kinases do not rely on phosphorylation for activation and new roles for unphosphorylated response regulators have been identified. The significance of recent findings and suggestions for further research are discussed.
© 2016 John Wiley & Sons Ltd.
Figures
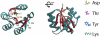

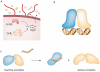
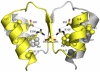

Similar articles
-
Atypical modes of bacterial histidine kinase signaling.Mol Microbiol. 2017 Jan;103(2):197-202. doi: 10.1111/mmi.13525. Epub 2016 Sep 30. Mol Microbiol. 2017. PMID: 27618209 Free PMC article. Review.
-
Bacterial hybrid histidine kinases in plant-bacteria interactions.Microbiology (Reading). 2016 Oct;162(10):1715-1734. doi: 10.1099/mic.0.000370. Epub 2016 Sep 8. Microbiology (Reading). 2016. PMID: 27609064 Review.
-
Evolution of orphan and atypical histidine kinases and response regulators for microbial signaling diversity.Int J Biol Macromol. 2024 Aug;275(Pt 1):133635. doi: 10.1016/j.ijbiomac.2024.133635. Epub 2024 Jul 2. Int J Biol Macromol. 2024. PMID: 38964677
-
Molecular Mechanisms of Two-Component Signal Transduction.J Mol Biol. 2016 Sep 25;428(19):3752-75. doi: 10.1016/j.jmb.2016.08.003. Epub 2016 Aug 9. J Mol Biol. 2016. PMID: 27519796 Free PMC article. Review.
-
The Histidine Residue of QseC Is Required for Canonical Signaling between QseB and PmrB in Uropathogenic Escherichia coli.J Bacteriol. 2017 Aug 22;199(18):e00060-17. doi: 10.1128/JB.00060-17. Print 2017 Sep 15. J Bacteriol. 2017. PMID: 28396353 Free PMC article.
Cited by
-
Proteolysis and multimerization regulate signaling along the two-component regulatory system AdeRS.iScience. 2021 Apr 26;24(5):102476. doi: 10.1016/j.isci.2021.102476. eCollection 2021 May 21. iScience. 2021. PMID: 34113820 Free PMC article.
-
Comparative genomics reveals an SNP potentially leading to phenotypic diversity of Salmonella enterica serovar Enteritidis.Microb Genom. 2021 May;7(5):000572. doi: 10.1099/mgen.0.000572. Microb Genom. 2021. PMID: 33952386 Free PMC article.
-
Single cell, super-resolution imaging reveals an acid pH-dependent conformational switch in SsrB regulates SPI-2.Elife. 2019 Apr 29;8:e45311. doi: 10.7554/eLife.45311. Elife. 2019. PMID: 31033442 Free PMC article.
-
Tetrachloroethene respiration in Sulfurospirillum species is regulated by a two-component system as unraveled by comparative genomics, transcriptomics, and regulator binding studies.Microbiologyopen. 2020 Dec;9(12):e1138. doi: 10.1002/mbo3.1138. Epub 2020 Nov 26. Microbiologyopen. 2020. PMID: 33242236 Free PMC article.
-
Regulation of Sporangium Formation, Spore Dormancy, and Sporangium Dehiscence by a Hybrid Sensor Histidine Kinase in Actinoplanes missouriensis: Relationship with the Global Transcriptional Regulator TcrA.J Bacteriol. 2020 Oct 8;202(21):e00228-20. doi: 10.1128/JB.00228-20. Print 2020 Oct 8. J Bacteriol. 2020. PMID: 32839172 Free PMC article.
References
-
- Baikalov I, Schroder I, Kaczor-Grzeskowiak M, Grzeskowiak K, Gunsalus RP, Dickerson RE. Structure of the Escherichia coli response regulator NarL. Biochemistry. 1996;35:11053–11061. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

