New insights into the Shwachman-Diamond Syndrome-related haematological disorder: hyper-activation of mTOR and STAT3 in leukocytes
- PMID: 27658964
- PMCID: PMC5034238
- DOI: 10.1038/srep33165
New insights into the Shwachman-Diamond Syndrome-related haematological disorder: hyper-activation of mTOR and STAT3 in leukocytes
Abstract
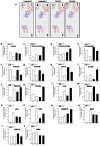
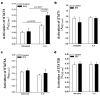
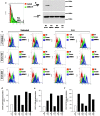
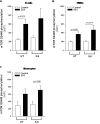
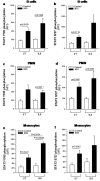
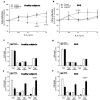
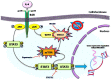
Shwachman-Diamond syndrome (SDS) is an inherited disease caused by mutations of a gene encoding for SBDS protein. So far little is known about SBDS exact function. SDS patients present several hematological disorders, including neutropenia and myelodysplastic syndrome (MDS), with increased risk of leukemic evolution. So far, the molecular mechanisms that underlie neutropenia, MDS and AML in SDS patients have been poorly investigated. STAT3 is a key regulator of several cellular processes including survival, differentiation and malignant transformation. Moreover, STAT3 has been reported to regulate neutrophil granulogenesis and to induce several kinds of leukemia and lymphoma. STAT3 activation is known to be regulated by mTOR, which in turn plays an important role in cellular growth and tumorigenesis. Here we show for the first time, to the best of our knowledge, that both EBV-immortalized B cells and primary leukocytes obtained from SDS patients present a constitutive hyper-activation of mTOR and STAT3 pathways. Interestingly, loss of SBDS expression is associated with this process. Importantly, rapamycin, a well-known mTOR inhibitor, is able to reduce STAT3 phosphorylation to basal levels in our experimental model. A novel therapeutic hypothesis targeting mTOR/STAT3 should represent a significant step forward into the SDS clinical practice.
Figures


References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

