Ovarian transcriptome associated with reproductive senescence in the long-living Ames dwarf mice
- PMID: 27663076
- PMCID: PMC5123904
- DOI: 10.1016/j.mce.2016.09.019
Ovarian transcriptome associated with reproductive senescence in the long-living Ames dwarf mice
Abstract
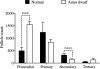
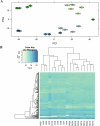
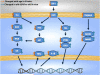
The aim of the current work was to evaluate the ovarian follicle reserve and the ovarian transcriptome in Ames dwarf (df/df) mice. The results suggest a delayed ovarian aging in df/df mice compared to normal (N) mice. Although a high number of genes were differentially expressed during aging of N mice, only a small fraction of these changed with aging in df/df mice. These alterations involved more than 500 categorized biological processes. The majority of these biological processes, including inflammatory/immune responses, were up-regulated with aging in N mice, while old df/df mice were characterized by down-regulation of these same processes in comparison to age matched N mice. However, biological processes related to DNA damage and repairing were commonly down-regulated with aging in both genotypes. In conclusion, delayed ovarian aging in long-living df/df mice was associated with reduced expression of genes related to the inflammatory and immune responses.
Keywords: GH; IGF; Ovarian aging; Transcriptome; mRNA.
Copyright © 2016 Elsevier Ireland Ltd. All rights reserved.
Figures


Similar articles
-
Ovarian aging and the activation of the primordial follicle reserve in the long-lived Ames dwarf and the short-lived bGH transgenic mice.Mol Cell Endocrinol. 2017 Nov 5;455:23-32. doi: 10.1016/j.mce.2016.10.015. Epub 2016 Oct 19. Mol Cell Endocrinol. 2017. PMID: 27771355 Free PMC article.
-
Changes of Ovarian microRNA Profile in Long-Living Ames Dwarf Mice during Aging.PLoS One. 2017 Jan 3;12(1):e0169213. doi: 10.1371/journal.pone.0169213. eCollection 2017. PLoS One. 2017. PMID: 28046124 Free PMC article.
-
Array-based expression analysis of mouse liver genes: effect of age and of the longevity mutant Prop1df.J Gerontol A Biol Sci Med Sci. 2001 Feb;56(2):B72-80. doi: 10.1093/gerona/56.2.b72. J Gerontol A Biol Sci Med Sci. 2001. PMID: 11213270
-
Dwarf mice as models for reproductive ageing research.Reprod Biomed Online. 2022 Jan;44(1):5-13. doi: 10.1016/j.rbmo.2021.09.016. Epub 2021 Oct 3. Reprod Biomed Online. 2022. PMID: 34794884 Review.
-
Calorie restriction and dwarf mice in gerontological research.Gerontology. 2010;56(4):404-9. doi: 10.1159/000235720. Epub 2009 Aug 19. Gerontology. 2010. PMID: 19690401 Review.
Cited by
-
Growth hormone-mediated reprogramming of macrophage transcriptome and effector functions.Sci Rep. 2019 Dec 18;9(1):19348. doi: 10.1038/s41598-019-56017-6. Sci Rep. 2019. PMID: 31852980 Free PMC article.
-
Growth Hormone Deficiency: Health and Longevity.Endocr Rev. 2019 Apr 1;40(2):575-601. doi: 10.1210/er.2018-00216. Endocr Rev. 2019. PMID: 30576428 Free PMC article. Review.
-
The role of cellular senescence in ovarian aging.NPJ Aging. 2024 Jul 20;10(1):35. doi: 10.1038/s41514-024-00157-1. NPJ Aging. 2024. PMID: 39033161 Free PMC article. Review.
-
Growth hormone increases DNA damage in ovarian follicles and macrophage infiltration in the ovaries.Geroscience. 2022 Apr;44(2):1071-1081. doi: 10.1007/s11357-021-00380-8. Epub 2021 May 5. Geroscience. 2022. PMID: 33954912 Free PMC article.
-
Growth hormone and aging.Rev Endocr Metab Disord. 2021 Mar;22(1):71-80. doi: 10.1007/s11154-020-09593-2. Epub 2020 Oct 1. Rev Endocr Metab Disord. 2021. PMID: 33001358 Review.
References
-
- Baker TG. A Quantitative and Cytological Study of Germ Cells in Human Ovaries. Proc R Soc Lond B Biol Sci. 1963;158:417–33. - PubMed
-
- Bartke A, Brown-Borg H, Mattison J, Kinney B, Hauck S, Wright C. Prolonged longevity of hypopituitary dwarf mice. Exp Gerontol. 2001;36:21–8. - PubMed
-
- Bartke A. Minireview: role of the growth hormone/insulin-like growth factor system in mammalian aging. Endocrinology. 2005;146:3718–23. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

