Global tRNA misacylation induced by anaerobiosis and antibiotic exposure broadly increases stress resistance in Escherichia coli
- PMID: 27672035
- PMCID: PMC5137444
- DOI: 10.1093/nar/gkw856
Global tRNA misacylation induced by anaerobiosis and antibiotic exposure broadly increases stress resistance in Escherichia coli
Abstract
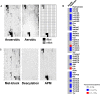
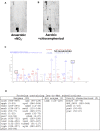
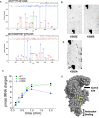
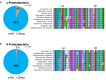
High translational fidelity is commonly considered a requirement for optimal cellular health and protein function. However, recent findings have shown that inducible mistranslation specifically with methionine engendered at the tRNA charging level occurs in mammalian cells, yeast and archaea, yet it was unknown whether bacteria were capable of mounting a similar response. Here, we demonstrate that Escherichia coli misacylates non-methionyl-tRNAs with methionine in response to anaerobiosis and antibiotic exposure via the methionyl-tRNA synthetase (MetRS). Two MetRS succinyl-lysine modifications independently confer high tRNA charging fidelity to the otherwise promiscuous, unmodified enzyme. Strains incapable of tRNA mismethionylation are less adept at growth in the presence of antibiotics and stressors. The presence of tRNA mismethionylation and its potential role in mistranslation within the bacterial domain establishes this response as a pervasive biological mechanism and connects it to diverse cellular functions and modes of fitness.
© The Author(s) 2016. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
Misacylation of tRNA with methionine in Saccharomyces cerevisiae.Nucleic Acids Res. 2012 Nov 1;40(20):10494-506. doi: 10.1093/nar/gks805. Epub 2012 Aug 31. Nucleic Acids Res. 2012. PMID: 22941646 Free PMC article.
-
Misacylation of specific nonmethionyl tRNAs by a bacterial methionyl-tRNA synthetase.Proc Natl Acad Sci U S A. 2011 Apr 26;108(17):6933-8. doi: 10.1073/pnas.1019033108. Epub 2011 Apr 11. Proc Natl Acad Sci U S A. 2011. PMID: 21482813 Free PMC article.
-
tRNA Misacylation with Methionine in the Mouse Gut Microbiome in Situ.Microb Ecol. 2017 Jul;74(1):10-14. doi: 10.1007/s00248-016-0928-0. Epub 2017 Jan 9. Microb Ecol. 2017. PMID: 28070678 Free PMC article.
-
Methionine as translation start signal: a review of the enzymes of the pathway in Escherichia coli.Biochimie. 1993;75(12):1061-75. doi: 10.1016/0300-9084(93)90005-d. Biochimie. 1993. PMID: 8199241 Review.
-
Methionyl-tRNA synthetase.Acta Biochim Pol. 2001;48(2):337-50. Acta Biochim Pol. 2001. PMID: 11732605 Review.
Cited by
-
A genetic polymorphism that is associated with mitochondrial energy metabolism increases risk of fibromyalgia.Pain. 2020 Dec;161(12):2860-2871. doi: 10.1097/j.pain.0000000000001996. Pain. 2020. PMID: 32658146 Free PMC article.
-
Genome-wide screening reveals metabolic regulation of stop-codon readthrough by cyclic AMP.Nucleic Acids Res. 2023 Oct 13;51(18):9905-9919. doi: 10.1093/nar/gkad725. Nucleic Acids Res. 2023. PMID: 37670559 Free PMC article.
-
Global mistranslation increases cell survival under stress in Escherichia coli.PLoS Genet. 2020 Mar 9;16(3):e1008654. doi: 10.1371/journal.pgen.1008654. eCollection 2020 Mar. PLoS Genet. 2020. PMID: 32150542 Free PMC article.
-
Increased mistranslation protects E. coli from protein misfolding stress due to activation of a RpoS-dependent heat shock response.FEBS Lett. 2019 Nov;593(22):3220-3227. doi: 10.1002/1873-3468.13578. Epub 2019 Aug 24. FEBS Lett. 2019. PMID: 31419308 Free PMC article.
-
Lysine Acetylation Regulates Alanyl-tRNA Synthetase Activity in Escherichia coli.Genes (Basel). 2018 Sep 28;9(10):473. doi: 10.3390/genes9100473. Genes (Basel). 2018. PMID: 30274179 Free PMC article.
References
-
- Reynolds N.M., Lazazzera B.A., Ibba M. Cellular mechanisms that control mistranslation. Nat. Rev. Microbiol. 2010;8:849–856. - PubMed
-
- Ibba M., Soll D. Quality control mechanisms during translation. Science. 1999;286:1893–1897. - PubMed
-
- Dale T., Uhlenbeck O.C. Amino acid specificity in translation. Trends Biochem. Sci. 2005;30:659–665. - PubMed
-
- Ibba M., Soll D. Aminoacyl-tRNA synthesis. Annu. Rev. Biochem. 2000;69:617–650. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

