A potential role for genome structure in the translation of mechanical force during immune cell development
- PMID: 27673560
- PMCID: PMC5120600
- DOI: 10.1080/19491034.2016.1238998
A potential role for genome structure in the translation of mechanical force during immune cell development
Abstract
Immune cells react to a wide range of environments, both chemical and physical. While the former has been extensively studied, there is growing evidence that physical and in particular mechanical forces also affect immune cell behavior and development. In order to elicit a response that affects immune cell behavior or development, environmental signals must often reach the nucleus. Chemical and mechanical signals can initiate signal transduction pathways, but mechanical forces may also have a more direct route to the nucleus, altering nuclear shape via mechanotransduction. The three-dimensional organization of DNA allows for the possibility that altering nuclear shape directly remodels chromatin, redistributing critical regulatory elements and proteins, and resulting in wide-scale gene expression changes. As such, integrating mechanotransduction and genome architecture into the immunology toolkit will improve our understanding of immune development and disease.
Keywords: Hi-C; chromatin; genome biology; immune; mechanosensory; mechanotransduction; nuclear lamin; nucleoskeleton; nucleus; tensegrity.
Figures
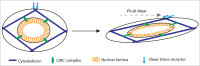


Similar articles
-
Mechanical Forces in Nuclear Organization.Cold Spring Harb Perspect Biol. 2022 Jan 4;14(1):a039685. doi: 10.1101/cshperspect.a039685. Cold Spring Harb Perspect Biol. 2022. PMID: 34187806 Free PMC article. Review.
-
Mechanotransduction and epigenetic modulations of chromatin: Role of mechanical signals in gene regulation.J Cell Biochem. 2024 Mar;125(3):e30531. doi: 10.1002/jcb.30531. Epub 2024 Feb 12. J Cell Biochem. 2024. PMID: 38345428 Review.
-
Nuclear shape, mechanics, and mechanotransduction.Circ Res. 2008 Jun 6;102(11):1307-18. doi: 10.1161/CIRCRESAHA.108.173989. Circ Res. 2008. PMID: 18535268 Free PMC article. Review.
-
LINC complex regulation of genome organization and function.Curr Opin Genet Dev. 2021 Apr;67:130-141. doi: 10.1016/j.gde.2020.12.007. Epub 2021 Jan 30. Curr Opin Genet Dev. 2021. PMID: 33524904 Review.
-
Dynamic regulation of nuclear architecture and mechanics-a rheostatic role for the nucleus in tailoring cellular mechanosensitivity.Nucleus. 2017 May 4;8(3):287-300. doi: 10.1080/19491034.2017.1285988. Epub 2017 Feb 2. Nucleus. 2017. PMID: 28152338 Free PMC article. Review.
Cited by
-
Graphene Oxide-Based Biocompatible 3D Mesh with a Tunable Porosity and Tensility for Cell Culture.ACS Biomater Sci Eng. 2018 May 14;4(5):1505-1517. doi: 10.1021/acsbiomaterials.8b00190. Epub 2018 Mar 29. ACS Biomater Sci Eng. 2018. PMID: 33445308 Free PMC article.
References
-
- Cohn M. A new concept of immune specificity emerges from a consideration of the self–nonself discrimination. Cell Immunol 1997; 181:103-8; PMID:9398397; http://dx.doi.org/ 10.1006/cimm.1997.1212 - DOI - PubMed
-
- Pelanda R, Piccirillo CA. Tolerance, immune regulation, and autoimmunity: cells and cytokines that make a difference. Curr Opin Immunol 2008; 20:629-31; PMID:18977298; http://dx.doi.org/ 10.1016/j.coi.2008.10.005 - DOI - PMC - PubMed
-
- Kochi Y, Suzuki A, Yamamoto K. Genetic basis of rheumatoid arthritis: A current review. Biochem Biophys Res Commun 2014; 452:254-62; PMID:25078624; http://dx.doi.org/ 10.1016/j.bbrc.2014.07.085 - DOI - PubMed
-
- Gregor MF, Hotamisligil GS. Inflammatory mechanisms in obesity. Annu Rev Immunol 2011; 29:415-45; PMID:21219177; http://dx.doi.org/ 10.1146/annurev-immunol-031210-101322 - DOI - PubMed
-
- Zhu J, Yamane H, Paul W. Differentiation of effector CD4 T cell populations. Annu Rev Immunol 2010; 28:445-89; PMID:20192806; http://dx.doi.org/ 10.1146/annurev-immunol-030409-101212 - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
