Mutations in Nonessential eIF3k and eIF3l Genes Confer Lifespan Extension and Enhanced Resistance to ER Stress in Caenorhabditis elegans
- PMID: 27690135
- PMCID: PMC5045169
- DOI: 10.1371/journal.pgen.1006326
Mutations in Nonessential eIF3k and eIF3l Genes Confer Lifespan Extension and Enhanced Resistance to ER Stress in Caenorhabditis elegans
Abstract
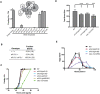
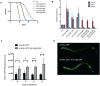
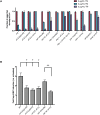
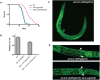
The translation initiation factor eIF3 is a multi-subunit protein complex that coordinates the assembly of the 43S pre-initiation complex in eukaryotes. Prior studies have demonstrated that not all subunits of eIF3 are essential for the initiation of translation, suggesting that some subunits may serve regulatory roles. Here, we show that loss-of-function mutations in the genes encoding the conserved eIF3k and eIF3l subunits of the translation initiation complex eIF3 result in a 40% extension in lifespan and enhanced resistance to endoplasmic reticulum (ER) stress in Caenorhabditis elegans. In contrast to previously described mutations in genes encoding translation initiation components that confer lifespan extension in C. elegans, loss-of-function mutations in eif-3.K or eif-3.L are viable, and mutants show normal rates of growth and development, and have wild-type levels of bulk protein synthesis. Lifespan extension resulting from EIF-3.K or EIF-3.L deficiency is suppressed by a mutation in the Forkhead family transcription factor DAF-16. Mutations in eif-3.K or eif-3.L also confer enhanced resistance to ER stress, independent of IRE-1-XBP-1, ATF-6, and PEK-1, and independent of DAF-16. Our data suggest a pivotal functional role for conserved eIF3k and eIF3l accessory subunits of eIF3 in the regulation of cellular and organismal responses to ER stress and aging.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Hansen M, Taubert S, Crawford D, Libina N, Lee S-J, Kenyon C. Lifespan extension by conditions that inhibit translation in Caenorhabditis elegans. Aging Cell 2007;6:95–110. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

