D3R grand challenge 2015: Evaluation of protein-ligand pose and affinity predictions
- PMID: 27696240
- PMCID: PMC5562487
- DOI: 10.1007/s10822-016-9946-8
D3R grand challenge 2015: Evaluation of protein-ligand pose and affinity predictions
Abstract
The Drug Design Data Resource (D3R) ran Grand Challenge 2015 between September 2015 and February 2016. Two targets served as the framework to test community docking and scoring methods: (1) HSP90, donated by AbbVie and the Community Structure Activity Resource (CSAR), and (2) MAP4K4, donated by Genentech. The challenges for both target datasets were conducted in two stages, with the first stage testing pose predictions and the capacity to rank compounds by affinity with minimal structural data; and the second stage testing methods for ranking compounds with knowledge of at least a subset of the ligand-protein poses. An additional sub-challenge provided small groups of chemically similar HSP90 compounds amenable to alchemical calculations of relative binding free energy. Unlike previous blinded Challenges, we did not provide cognate receptors or receptors prepared with hydrogens and likewise did not require a specified crystal structure to be used for pose or affinity prediction in Stage 1. Given the freedom to select from over 200 crystal structures of HSP90 in the PDB, participants employed workflows that tested not only core docking and scoring technologies, but also methods for addressing water-mediated ligand-protein interactions, binding pocket flexibility, and the optimal selection of protein structures for use in docking calculations. Nearly 40 participating groups submitted over 350 prediction sets for Grand Challenge 2015. This overview describes the datasets and the organization of the challenge components, summarizes the results across all submitted predictions, and considers broad conclusions that may be drawn from this collaborative community endeavor.
Keywords: D3R; Docking; Free energy; Ligand; Protein; Scoring.
Figures



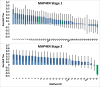

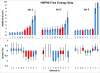
References
-
- Solit DB, Rosen N. Hsp90: a novel target for cancer therapy. Curr Top Med Chem. 2006;6:1205–1214. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

