Trans-ethnic meta-analysis of genome-wide association studies for Hirschsprung disease
- PMID: 27702942
- PMCID: PMC6078638
- DOI: 10.1093/hmg/ddw333
Trans-ethnic meta-analysis of genome-wide association studies for Hirschsprung disease
Abstract
Hirschsprung disease (HSCR) is the most common cause of neonatal intestinal obstruction. It is characterized by the absence of ganglia in the nerve plexuses of the lower gastrointestinal tract. So far, three common disease-susceptibility variants at the RET, SEMA3 and NRG1 loci have been detected through genome-wide association studies (GWAS) in Europeans and Asians to understand its genetic etiologies. Here we present a trans-ethnic meta-analysis of 507 HSCR cases and 1191 controls, combining all published GWAS results on HSCR to fine-map these loci and narrow down the putatively causal variants to 99% credible sets. We also demonstrate that the effects of RET and NRG1 are universal across European and Asian ancestries. In contrast, we detected a European-specific association of a low-frequency variant, rs80227144, in SEMA3 [odds ratio (OR) = 5.2, P = 4.7 × 10-10]. Conditional analyses on the lead SNPs revealed a secondary association signal, corresponding to an Asian-specific, low-frequency missense variant encoding RET p.Asp489Asn (rs9282834, conditional OR = 20.3, conditional P = 4.1 × 10-14). When in trans with the RET intron 1 enhancer risk allele, rs9282834 increases the risk of HSCR from 1.1 to 26.7. Overall, our study provides further insights into the genetic architecture of HSCR and has profound implications for future study designs.
© The Author 2016. Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Figures

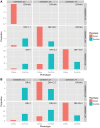
Similar articles
-
Population variation in total genetic risk of Hirschsprung disease from common RET, SEMA3 and NRG1 susceptibility polymorphisms.Hum Mol Genet. 2015 May 15;24(10):2997-3003. doi: 10.1093/hmg/ddv051. Epub 2015 Feb 9. Hum Mol Genet. 2015. PMID: 25666438 Free PMC article.
-
Association of genetic polymorphisms in the RET-protooncogene and NRG1 with Hirschsprung disease in Thai patients.J Hum Genet. 2012 May;57(5):286-93. doi: 10.1038/jhg.2012.18. Epub 2012 Mar 1. J Hum Genet. 2012. PMID: 22377709
-
Cumulative Risk Impact of RET, SEMA3, and NRG1 Polymorphisms Associated With Hirschsprung Disease in Han Chinese.J Pediatr Gastroenterol Nutr. 2017 Mar;64(3):385-390. doi: 10.1097/MPG.0000000000001263. J Pediatr Gastroenterol Nutr. 2017. PMID: 27203398
-
Association of REarranged during Transfection (RET) c.73 + 9277T > C and c.135G > a Polymorphisms with Susceptibility to Hirschsprung Disease: A Systematic Review and Meta-Analysis.Fetal Pediatr Pathol. 2020 Dec;39(6):476-490. doi: 10.1080/15513815.2019.1672225. Epub 2019 Oct 7. Fetal Pediatr Pathol. 2020. PMID: 31590591
-
Molecular mechanisms of RET-induced Hirschsprung pathogenesis.Ann Med. 2006;38(1):11-9. doi: 10.1080/07853890500442758. Ann Med. 2006. PMID: 16448984 Review.
Cited by
-
Genome-wide association study of Hirschsprung disease detects a novel low-frequency variant at the RET locus.Eur J Hum Genet. 2018 Apr;26(4):561-569. doi: 10.1038/s41431-017-0053-7. Epub 2018 Jan 29. Eur J Hum Genet. 2018. PMID: 29379196 Free PMC article.
-
Association of Variants in PLD1, 3p24.1, and 10q11.21 Regions With Hirschsprung's Disease in Han Chinese Population.Front Genet. 2020 Jul 10;11:738. doi: 10.3389/fgene.2020.00738. eCollection 2020. Front Genet. 2020. PMID: 32765588 Free PMC article.
-
Association of rs2435357 and rs2506030 polymorphisms in RET with susceptibility to hirschsprung disease: A systematic review and meta-analysis.Front Pediatr. 2022 Oct 17;10:1030933. doi: 10.3389/fped.2022.1030933. eCollection 2022. Front Pediatr. 2022. PMID: 36324815 Free PMC article. Review.
-
Goldberg-Shprintzen syndrome is determined by the absence, or reduced expression levels, of KIFBP.Hum Mutat. 2020 Nov;41(11):1906-1917. doi: 10.1002/humu.24097. Epub 2020 Sep 16. Hum Mutat. 2020. PMID: 32939943 Free PMC article.
-
A gene regulatory network explains RET-EDNRB epistasis in Hirschsprung disease.Hum Mol Genet. 2019 Sep 15;28(18):3137-3147. doi: 10.1093/hmg/ddz149. Hum Mol Genet. 2019. PMID: 31313802 Free PMC article.
References
-
- Amiel J., Sproat-Emison E., Garcia-Barcelo M., Lantieri F., Burzynski G., Borrego S., Pelet A., Arnold S., Miao X., Griseri P., et al. (2008) Hirschsprung disease, associated syndromes and genetics: a review. J. Med. Genet., 45, 1–14. - PubMed
-
- Tam P.K., Garcia-Barcelo M. (2009) Genetic basis of Hirschsprung's disease. Pediatr. Surg. Int., 25, 543–558. - PubMed
-
- Chakravarti A., McCallion A.S., Lyonnet S. (2014) In Beaudet A.L., Vogelstein B., Kinzler K.W., Antonarakis S.E., Ballabio A., Gibson K.M., Mitchell G. (eds), The Online Metabolic and Molecular Bases of Inherited Disease. The McGraw-Hill Companies, Inc, New York, NY.
-
- Alves M.M., Sribudiani Y., Brouwer R.W., Amiel J., Antinolo G., Borrego S., Ceccherini I., Chakravarti A., Fernandez R.M., Garcia-Barcelo M.M., et al. (2013) Contribution of rare and common variants determine complex diseases-Hirschsprung disease as a model. Dev. Biol, 382, 320–329. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

