Linear ubiquitination by LUBEL has a role in Drosophila heat stress response
- PMID: 27702987
- PMCID: PMC5090701
- DOI: 10.15252/embr.201642378
Linear ubiquitination by LUBEL has a role in Drosophila heat stress response
Abstract
The HOIP ubiquitin E3 ligase generates linear ubiquitin chains by forming a complex with HOIL-1L and SHARPIN in mammals. Here, we provide the first evidence of linear ubiquitination induced by a HOIP orthologue in Drosophila We identify Drosophila CG11321, which we named Linear Ubiquitin E3 ligase (LUBEL), and find that it catalyzes linear ubiquitination in vitro We detect endogenous linear ubiquitin chain-derived peptides by mass spectrometry in Drosophila Schneider 2 cells and adult flies. Furthermore, using CRISPR/Cas9 technology, we establish linear ubiquitination-defective flies by mutating residues essential for the catalytic activity of LUBEL Linear ubiquitination signals accumulate upon heat shock in flies. Interestingly, flies with LUBEL mutations display reduced survival and climbing defects upon heat shock, which is also observed upon specific LUBEL depletion in muscle. Thus, LUBEL is involved in the heat response by controlling linear ubiquitination in flies.
Keywords: LUBEL; deubiquitinase; linear chain; ubiquitin; ubiquitin E3 ligase.
© 2016 The Authors. Published under the terms of the CC BY 4.0 license.
Figures

- A
Schematics of eight different linkage types of Ub chains. Ub can form different types of polymers by conjugating via intrinsic Lys residues (K 6, K 11, K 27, K 29, K 33, K 48, and K 63) and M 1 (linear).
- B
A multiple amino acid sequence alignment of RBR‐C in HOIP family members. Conserved residues are colored according to the ClustalX coloring scheme. Long unaligned regions in A. mellifera and P. humanus were replaced by the number of deleted residues in squared brackets. R1, IBR, R2, and LDD domains are indicated with gray bars above the sequences. The zinc (Zn)‐coordinating residues known in H. sapiens HOIP‐RBR‐C are labeled below. A black triangle indicates the Cys (C) residue for the thioester intermediate, two * are Zn‐coordinating Cys residues in R2 targeted to create catalytically dead mutants, and two O are residues mutated in LDD.
- C–F
Multiple amino acid sequence alignments of N‐terminal HOIP domains, UBA1 (C), UBA2 (D), B‐box (E), and NZF (F).

Schematic diagram of HOIP orthologues in Drosophila melanogaster (D. melanogaster) and Homo sapiens (H. sapiens). Drosophila melanogaster CG11321 is predicted to contain B‐box (blue), Npl4 zinc finger (NZF) (navy), ubiquitin (Ub)‐associated (UBA1 and UBA2) (pink), and a C‐terminal catalytic region, RING between RING (RBR)‐C. The RBR‐C region consists of RING1 (R1; red), in‐between RING (IBR; yellow), RING2 (R2; light green), and linear Ub chain determining domain (LDD; green). PNGase/Ub‐associated (PUB) (sky blue) and zinc finger (ZF) (lavender) domains were found in H. sapiens but not in D. melanogaster.
Ub chain interaction of the LUBEL‐NZF and UBA2 domains. Immobilized GST‐NZF or GST‐UBA2 was incubated with Lys (K) 48‐, K 63‐linked, or linear di‐Ub (Ub2) chains and bound Ub chains were subjected to immunoblotting using anti‐Ub antibody or a linear linkage‐specific Ub antibody (anti‐linear Ub antibody). Loading of GST‐fusion proteins was visualized by Ponceau S staining.

In vitro ubiquitination assays of LUBEL‐RBR‐C WT or catalytically dead C2704A in combination with Ube1 and two different Drosophila E2s, UbcD10, or Effete/UbcD1. Reactions were terminated at indicated times, and synthesized Ub chains were detected by immunoblotting using anti‐linear Ub antibody. Total protein loading was visualized by Ponceau S staining. *: nonspecific band.
In vitro deubiquitination of Ub chains synthesized by HOIP‐ and LUBEL‐RBR‐C. Ub chains produced by RBR‐C (human or Drosophila) with UbcH7, UbcD10 or Effete were incubated with a linear linkage‐specific DUB, OTULIN (WT or catalytically dead C129A (C/A) mutant), or a Lys‐linkage‐specific DUB, vOTU. The DUB‐treated samples were subjected to Coomassie staining or immunoblotting using anti‐Ub antibody. *: nonspecific band.
Thioester formation assay using Atto647‐labeled Ub. Ubiquitination assay using LUBEL‐RBR‐C WT (left panel) or C2704A (right panel): lanes 1/2 Atto‐Ub + E1, lanes 3/4 + ATP, lanes 5/6 + E2, lanes 7/8 + E3, and lanes 9/10 + Ub‐His6. In the presence of N‐terminally tagged Atto‐Ub and C‐terminally tagged Ub‐His6, only Ub2 product can be synthesized and longer chain formation is restricted. Samples are run without or with DTT in odd or even numbered lanes, respectively. The gels monitor the Atto‐labeled Ub.
The linear Ub chain formation activity of LUBEL‐LDD mutants. Recombinant proteins of RBR‐C WT, C2704A mutant, and LDD mutants (R2754A and D2755A) were assessed for their activity by in vitro ubiquitination assay. The samples were resolved on a gel and stained with Coomassie or immunoblotted using anti‐linear Ub antibody.
Linear Ub chain formation by full‐length LUBEL transient expression in insect cells. Full‐length Myc‐LUBEL was transiently expressed in Drosophila Schneider 2 (S2) cells, and Myc‐HOIP alone or with HA‐HOIL‐1L in HEK293T cells. Total cell lysates (TCL) of control and transfected samples were incubated with immobilized GST‐Linear‐Tandem Ub binding entity (Linear‐TUBE) containing three tandem repeats of ABIN‐1‐UBAN. Pulldown samples were blotted with anti‐linear Ub antibody, while TCL were blotted with anti‐Myc antibody for exogenous LUBEL and HOIP, anti‐HA antibody for HOIL‐1L, and anti‐tubulin antibody for loading. Input of GST proteins was analyzed by Ponceau S staining. *: nonspecific band.

- A
In vitro ubiquitination assay of predicted LUBEL‐RBR‐C in combination with Ube1 and two different Drosophila E2s, UbcD10 and Effete/UbcD1, using nontagged or N‐terminally His6‐tagged Ub. Synthesized Ub chains were analyzed by immunoblotting using anti‐linear Ub antibody or anti‐Ub antibody. Protein loading was visualized by Ponceau S staining. *: nonspecific band.
- B
Mass spectrometry analysis of the Ub chains generated by LUBEL‐RBR‐C and UbcD10. MS/MS spectra acquired from the linear Ub chain peptide using an identical sample as Fig 2A lane 4 is shown.
- C, D
In vitro deubiquitinating activities of vOTU (C), or OTULIN (D). Recombinant vOTU, OTULIN (WT or a catalytically dead C129A mutant) was incubated with K 48‐, K 63‐linked, or linear Ub2 chains for indicated times. Subsequently, proteins were resolved on SDS–PAGE gels and stained with Coomassie dye. *: nonspecific band.
- E
In vitro ubiquitination assay of LUBEL‐RBR‐C C2690S/C2693S (CC/SS) mutant compared to WT. Linear Ub chain formation was analyzed by immunoblotting using anti‐linear Ub antibody. Total amount of proteins was analyzed by Ponceau S staining. *: nonspecific band.
- F
Linear Ub chains in Drosophila S2 cells transfected with LUBEL‐RBR‐C. Myc‐RBR‐C WT or Myc‐RBR‐C CC/SS was transfected in S2 cells. Linear Ub chains in TCL were visualized by immunoblotting using anti‐linear Ub antibody. Expression of RBR‐C (WT or C2690/2693S) was analyzed by using anti‐Myc‐antibody, and tubulin was blotted to examine protein loading.
- G
In vitro ubiquitination assay using full‐length LUBEL. In vitro ubiquitination assay was performed using recombinant full‐length LUBEL purified by a baculovirus‐based insect expression method, in combination with Ube1 and UbcD10. Amount of E3 ligase was determined by using an antibody raised against LUBEL‐RBR‐C. LUBEL‐RBR‐C was used as positive control.
- H
Catalytic activity of LUBEL‐RBR‐C with extended N‐terminal UBA2 in vitro. In vitro ubiquitination assay was performed using LUBEL‐RBR‐C or LUBEL‐UBA2‐RBR‐C in combination with Ube1 and UbcD10. Linear Ub chain formation was analyzed as (E), and amount of E3 ligase was determined by anti‐RBR‐C antibody. *: nonspecific band.
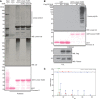
Endogenous linear Ub chains enriched in total protein extracts (input, lane 1) from adult w1118 (w −) flies by GST‐Linear‐TUBE (lane 3). The enriched samples were further treated with recombinant DUBs, vOTU (lane 4), or OTULIN (lane 5). Ub chains were visualized by immunoblotting using anti‐linear Ub antibody or anti‐Ub antibody. Loading of GST proteins was analyzed by Ponceau S staining. *: nonspecific band.
Endogenous linear Ub chains in total cell lysate (TCL) of S2 cells expressing human Flag‐OTULIN WT or catalytically inactive C129A mutant. TCLs enriched with GST‐Linear‐TUBE were examined by immunoblotting using anti‐linear Ub antibody. Expression of OTULIN was analyzed by anti‐Flag antibody, and loading of TCL was detected by anti‐tubulin antibody. Input of GST proteins was analyzed by Ponceau S staining. *: nonspecific band.
Mass spectrometry analysis using an identical sample as in panel (B) lane 6. MS/MS spectra of the prototypic linear Ub chain peptide (GGMQIFVK) is shown.
- A, B
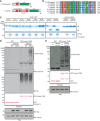
Schematic diagram of Drosophila (D. melanogaster) CYLD (dCYLD) and human (H. sapiens) CYLD (A), and a multiple amino acid sequence alignment of a catalytic region of the ubiquitin carboxyl‐terminal hydrolase (UCH) domain in different species (B). dCYLD is 639 amino acids in length, and contains one cytoskeletal‐associated protein–glycine‐conserved (CAP‐Gly) domain (Red) and the UCH domain (Green) (A). Human CYLD is 965 amino acids in length and contains three CAP‐Gly domains, an unstructured phosphorylation region (Phos‐site (Pink); specific for CYLD chordata species), and a UCH domain (Green). A conserved active Cys (C284 in Drosophila) is indicated by * (B).
- C
In vitro deubiquitination assay of dCYLD. dCYLD was incubated with all eight linkage types of Ub2 chains. dCYLD catalytically dead C284S mutant was incubated with K 63‐ or linear Ub2 chains. All the proteins in the reactions were resolved on Coomassie‐stained SDS–PAGE gel. *: nonspecific band.
- D
Stabilization of linear Ub chains by dCYLD C284S mutant transient expression in S2 cells. Myc‐tagged dCYLD WT and C284S were transfected in S2 cells, and linear Ub chains were enriched with GST‐Linear‐TUBE and immunoblotted by anti‐linear Ub or anti‐Ub antibodies. Input of GST proteins was visualized by Ponceau S staining. Expression of Myc‐dCYLD was examined by anti‐Myc antibody, and anti‐tubulin antibody was used for the loading control of TCL. *: nonspecific band.
- E
Linear Ub chains in dCYLD mutant flies. Endogenous levels of linear Ub chains in dCYLD mutant flies were compared with a control w − fly strain by performing GST‐Linear‐TUBE pulldown. Enriched linear Ub chains were immunoblotted with anti‐linear Ub antibody. Input of GST proteins was visualized by Ponceau S staining. Tubulin was used for the loading control of the input. *: nonspecific band.

A multiple amino acid sequence alignment of the CYLD catalytic domain in different species. * indicates predicted catalytic Cys residue, while C‐X‐X‐C pairs (labeled CxxC) and the UCH catalytic domains, UCH2‐1 and UCH2‐2 (in gray), are shown below the sequences.
Endogenous level of K 63‐linked Ub chains in dCYLD mutant flies. Poly‐Ub chains in the total protein extracts of w − and dCYLD mutant were enriched by GST‐TR‐TUBE pulldown, and the samples were resolved and detected using anti‐K 63‐linked Ub chains. GST was used as control for the pulldown. Input of GST proteins was visualized by Ponceau S, and total protein extracts were blotted with anti‐tubulin antibody. *: nonspecific band.
Interaction between dCYLD and LUBEL‐RBR‐C. Flag‐RBR‐C was transfected into S2 cells and total cell lysate was incubated with either agarose‐immobilized GST or GST‐dCYLD. After GST pulldown, samples were analyzed by immunoblotting using anti‐Flag antibody. Loading of GST proteins was visualized by Ponceau S staining.
Protein–protein interaction of recombinant dCYLD and recombinant LUBEL‐RBR‐C. LUBEL‐RBR‐C purified from E. coli was incubated with immobilized GST or GST‐dCYLD for pulldown assay. The interaction was analyzed by immunoblotting using anti‐LUBEL‐RBR antibody. Loading of GST proteins was visualized by Ponceau S staining. *: nonspecific band.
- A
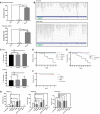
Endogenous mRNA expression of LUBEL detected in different embryonic stages of w − flies by qPCR. RNA was isolated from embryos and the LUBEL expression levels were measured by qPCR. TATA binding protein (TBP) was used as a reference, and two sets of primers, N‐terminal region or catalytic region, were used to detect all isoforms or only the long isoforms, respectively. Representative data are shown from three independent experiments.
- B
Endogenous mRNA fragments of LUBEL detected in S2 cells (above) and w − male adult flies (bottom) by RNA‐Seq. The aligned transcripts were visualized using The Integrative Genomics Viewer (IGV_2.3.40 software) and screenshots of the CG11321 region are shown. Representative alignments of three repeats are shown.
- C
Negative geotaxis assay of aged male (top panel) and female (bottom panel) flies. The assays were repeated five times for each group (between 47 to 55 flies per group), and the results were combined to create the graphs. t‐test analysis showed no significant difference between the lines. Representative videos can be found in Movies EV2 and EV3.
- D, E
Survival of LUBEL mutant and dCYLD mutant flies upon Gram‐negative bacterial infection. Septic injury was performed using E. coli in 20 young adult male flies. Recovered flies were kept in 25°C and counted every 24 h until indicated time. Rel E20 was used as positive control for the assay. Curve comparison tests indicated there is no statistical difference except for Rel E20 > 0.0001. Representative survival curve is shown for the LUBEL mutant flies, CC/SS #1 and #2, and delR2 (D; from four independent experiments), and catalytically dead dCYLD flies (E; from three independent experiments).
- F
Survival of adult male LUBEL mutant and dCLYD mutant flies by septic injury with Gram‐positive M. luteus bacteria. Septic injury with M. luteus was performed and recovered flies were kept in 29°C and counted every 24 h for indicated time. Curve comparison tests indicated the difference is not significant between the lines. Representative data are shown from three independent experiments.
- G
Antimicrobial peptide induction upon septic injury in LUBEL mutant flies. Septic injury with E. coli was performed, and after 10‐h recovery, RNA was isolated and mRNA levels of Attacin C, Diptericin, and Drosomycin were measured by qPCR. Rp49 was used as a reference, and w − unpricked sample was used as calibrator to calculate the expression ratio. Multiple comparisons were performed using one‐way ANOVA. Representative data are shown from three independent experiments.
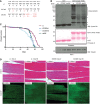
Amino acid sequences of the mutations introduced in LUBEL by CRISPR/Cas9. Two LUBEL mutant lines were created for in vivo studies; w − parental line, C2690S/C2693S (CC/SS), and delR2, which has an indel mutation with an early stop codon at amino acid 2,719 due to a frameshift. Mutations are indicated in red.
Linear Ub chains in w − and LUBEL mutant fly lysates. w −, CC/SS, and delR2 flies were analyzed for the level of endogenous linear Ub chains by enriching the linear ubiquitin chains by GST‐Linear‐TUBE. Pulldown samples were immunoblotted by anti‐linear Ub antibody, while GST proteins were analyzed by Ponceau S staining. Anti‐tubulin antibody was used for the loading control of the input. *: nonspecific band.
The life spans of female w − and catalytically dead LUBEL mutant flies (two clones of CC/SS, indicated as #1 and #2, and delR2). Survival of three independent cohorts with ˜80 flies each was monitored over time. Median survival time (days): w − = 78, CC/SS #1 = 68, CC/SS #2 = 65, and delR2 = 71. Total sample sizes are as follows: w − = 215, CC/SS #1 = 242, CC/SS #2 = 233, delR2 = 273. Log‐rank (Mantel–Cox) test between fly lines: w − and CC/SS #1 > 0.0001, w − and CC/SS #2 > 0.0001, w − and delR2 > 0.0001, CC/SS #1 and CC/SS #2 = 0.0007, CC/SS #1 and delR2 = not significant, CC/SS #2 and delR2 = not significant.
Histological analysis of muscle in LUBEL mutant flies. Hematoxylin and eosin (H&E) (top panels) and actin immunofluorescent staining (actin‐IF) (bottom panels) of thorax muscles from young (day 3) or aged (day 60) w − or CC/SS female flies. Scale bars: 50 μm.

The level of poly‐Ub chains in w − flies after heat shock. w − male flies were incubated in a 36°C water bath for indicated times and the levels of poly‐Ub chains in total protein extracts were compared by immunoblotting using antibodies against linear, K 48‐, and K 63‐linked Ub chains. Ponceau S staining was used to visualize the protein loading.
Survival of catalytically dead LUBEL mutant flies upon heat shock. Young adult flies (15 males and 15 females) were incubated in a 36°C water bath and immobilized flies were counted over the time indicated. Median survival time (h): w − = 9.25, CC/SS #1 = 5.5, CC/SS #2 = 5, delR2 = 5.75, CC/SS #1/+ = 8.5, CC/SS #2/+ = 10, delR2/+ = 8. P‐values calculated by Gehan–Breslow–Wilcoxon test between fly lines: w − and CC/SS #1, CC/SS#2, or delR2 > 0.0001 (****P < 0.0001), w − and CC/SS #1/+ = 0.0134, w − and CC/SS #2/+ = ns, w − and delR2/+ = 0.0002, CC/SS #1 and CC/SS #1/+ > 0.0001, CC/SS #2 and CC/SS #2/+ > 0.0001, delR2 and delR2/+ > 0.0001. Representative data are shown from six independent experiments.
Survival of whole‐body or muscle‐specific LUBEL knockdown flies upon heat shock. shRNA‐based knockdown (KD) of LUBEL were driven by Tub‐Gal4 or Mef2‐Gal4 flies. Control fly lines (Tub‐Gal4/+ and Mef2‐Gal4/+) were used to compare with the KD flies. 15 males and 15 females per each line were used in this assay. Median survival time (h): Tub‐Gal4/+ = 7, Tub‐Gal4 > LUBEL = 3.5, Mef2‐Gal4/+ = 7.5, Mef2‐Gal4 > LUBEL = 7. P‐values calculated by Gehan–Breslow–Wilcoxon test: Tub‐Gal4 < 0.0001 (****), Mef2‐Gal4 = 0.0029 (**). Representative data are shown from three independent experiments.
Heat‐induced mRNA expression of HSP70. w −, CC/SS #2, and delR2 flies were heat treated for 60 min and recovered for 1 h, and mRNA HSP70 was quantified by qPCR. Rp49 was used as a reference, and w − untreated sample was used as calibrator to calculate the expression ratio. Data are analyzed by two‐way ANOVA with multiple comparison, and presented as mean ± SD (*P < 0.01, ***P < 0.001, ****P < 0.0001). Representative data are shown from three independent experiments.

Immunoblotting for total Ub chains in heat‐shocked w − flies in Fig 6A using anti‐pan Ub antibody.
mRNA expression of poly‐ubiquitin gene ubi‐p63E in LUBEL mutant flies, untreated, or heat shocked. 10 male and 10 female adult flies were heat shocked for 1 h and RNA was isolated and Ubi‐p63E mRNA level was measured by qPCR. Rp49 was used as reference and untreated samples were used as calibrator for each fly line to calculate the expression ratio. Data are analyzed by two‐way ANOVA with multiple comparison, and presented as mean ± SD (****P < 0.001). Representative of three experiments is shown.
Expression of LUBEL mRNA in heterozygous flies (Tub‐Gal4/+ and Mef2‐Gal4/+) and UAS‐shLUBEL (Tub‐Gal4 > LUBEL and Mef2‐Gal4 > LUBEL) flies detected by qPCR. The primers that target N‐terminal region (left panel) or catalytic region (right panel) of LUBEL were used. Rp49 was used as reference, and GD control fly line was used as calibrator to calculate the expression ratio. Data were analyzed using unpaired t‐test; values represent mean ± SD (**P < 0.01, ***P < 0.001), (n = 3). Representative of three independent experiments are shown.
Survival of dCYLD mutant flies upon heat shock. Survival curve of heat‐treated w − and dCYLD mutant flies is shown. 15 male and 15 female files per each fly line were used in this assay. Curve comparison tests indicated the difference is not significant. Representative data are shown from four independent experiments.
Heat‐induced mRNA expression of HSP70. w −, CC/SS #2, and delR2 flies were heat treated for 30 min and mRNA HSP70 was quantified by qPCR. Rp49 was used as a reference and w − untreated sample was used as calibrator to calculate the expression ratio. Data are analyzed by two‐way ANOVA with multiple comparison, and presented as mean ± SD (****P < 0.0001). Representative of three experiments is shown.
Repeat of muscle‐specific LUBEL KD using B24‐Gal4 driver. A heat‐hock survival assay was performed as Fig 6C, using 24B‐Gal4 driver line to knockdown LUBEL in the muscle. P‐values calculated by Gehan–Breslow–Wilcoxon test: Tub‐Gal4 < 0.0001 (****), 24B‐Gal4 = 0.0066 (**). knockdown efficiency for the catalytic region, analyzed as in (C), is shown on the right graph. Representative of three experiments is shown.
References
-
- Sasaki K, Iwai K (2015) Roles of linear ubiquitinylation, a crucial regulator of NF‐kappaB and cell death, in the immune system. Immunol Rev 266: 175–189 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

