Deregulation of the pRb-E2F4 axis alters epidermal homeostasis and favors tumor development
- PMID: 27708231
- PMCID: PMC5342772
- DOI: 10.18632/oncotarget.12362
Deregulation of the pRb-E2F4 axis alters epidermal homeostasis and favors tumor development
Erratum in
-
Correction: Deregulation of the pRb-E2F4 axis alters epidermal homeostasis and favors tumor development.Oncotarget. 2017 Jul 25;8(30):50324. doi: 10.18632/oncotarget.19511. Oncotarget. 2017. PMID: 28772360 Free PMC article. No abstract available.
Abstract
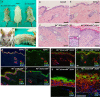
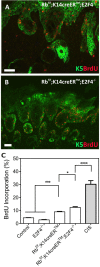
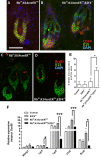
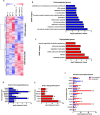
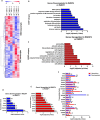
E2F/RB activity is altered in most human tumors. The retinoblastoma family of proteins plays a key role in regulating the progression of the cell cycle from the G1 to S phases. This is achieved through negative regulation of E2F transcription factors, important positive regulators of cell cycle entry. E2F family members are divided into two groups: activators (E2F1-E2F3a) and repressors (E2F3b-E2F8). E2F4 accounts for a large part of the E2F activity and is a main E2F repressor member in vivo. Perturbations in the balance from quiescence towards proliferation contribute to increased mitotic gene expression levels frequently observed in cancer. We have previously reported that combined Rb1-Rbl1 or Rb1-E2f1 ablation in epidermis produces important alterations in epidermal proliferation and differentiation, leading to tumor development. However, the possible roles of E2F4 in this context are still to be determined. Here, we show the absence of any discernible phenotype in the skin of mice lacking of E2f4. In contrast, the inducible loss of Rb1 in the epidermis of E2F4-null mice produced multiple skin abnormalities including altered differentiation and proliferation, spontaneous wounds, carcinoma in situ development and stem cell perturbations. All these phenotypic alterations are associated with extensive gene expression changes, the induction of c-myc and the Akt activation. Moreover the whole transcriptome analyses in comparison with previous models generated also revealed extensive changes in multiple repressive complexes and in transcription factor activity. These results point to E2F4 as a master regulator in multiple steps of epidermal homeostasis in Rb1 absence.
Keywords: E2F4; Rb; cancer; epidermis; mouse.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



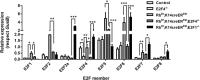
Similar articles
-
E2F1 loss induces spontaneous tumour development in Rb-deficient epidermis.Oncogene. 2013 Jun 13;32(24):2937-51. doi: 10.1038/onc.2012.316. Epub 2012 Aug 13. Oncogene. 2013. PMID: 22890321
-
Distinct capacities of individual E2Fs to induce cell cycle re-entry in postmitotic lens fiber cells of transgenic mice.Dev Neurosci. 2004;26(5-6):435-45. doi: 10.1159/000082285. Dev Neurosci. 2004. PMID: 15855772
-
TGFbeta-mediated formation of pRb-E2F complexes in human myeloid leukemia cells.Biochem Biophys Res Commun. 2008 May 2;369(2):277-80. doi: 10.1016/j.bbrc.2008.02.051. Epub 2008 Feb 21. Biochem Biophys Res Commun. 2008. PMID: 18294958 Review.
-
The ancient function of RB-E2F pathway: insights from its evolutionary history.Biol Direct. 2010 Sep 20;5:55. doi: 10.1186/1745-6150-5-55. Biol Direct. 2010. PMID: 20849664 Free PMC article.
-
The Rb family connects with the Tp53 family in skin carcinogenesis.Mol Carcinog. 2007 Aug;46(8):618-23. doi: 10.1002/mc.20338. Mol Carcinog. 2007. PMID: 17486638 Review.
Cited by
-
The Role of p16INK4a Pathway in Human Epidermal Stem Cell Self-Renewal, Aging and Cancer.Int J Mol Sci. 2017 Jul 22;18(7):1591. doi: 10.3390/ijms18071591. Int J Mol Sci. 2017. PMID: 28737694 Free PMC article. Review.
-
Identification of key genes and pathways for esophageal squamous cell carcinoma by bioinformatics analysis.Exp Ther Med. 2018 Aug;16(2):1121-1130. doi: 10.3892/etm.2018.6316. Epub 2018 Jun 15. Exp Ther Med. 2018. PMID: 30112053 Free PMC article.
References
-
- Frolov MV, Dyson NJ. Molecular mechanisms of E2F-dependent activation and pRB-mediated repression. J Cell Sci. 2004;117(Pt 11):2173–2181. - PubMed
-
- Paramio JM, Lain S, Segrelles C, Lane EB, Jorcano JL. Differential expression and functionally co-operative roles for the retinoblastoma family of proteins in epidermal differentiation. Oncogene. 1998;17:949–957. - PubMed
-
- Paramio JM, Segrelles C, Casanova ML, Jorcano JL. Opposite functions for E2F1 and E2F4 in human epidermal keratinocyte differentiation. J Biol Chem. 2000;275:41219–41226. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

