UV-Induced DNA Damage and Mutagenesis in Chromatin
- PMID: 27716995
- PMCID: PMC5315636
- DOI: 10.1111/php.12646
UV-Induced DNA Damage and Mutagenesis in Chromatin
Abstract
UV radiation induces photolesions that distort the DNA double helix and, if not repaired, can cause severe biological consequences, including mutagenesis or cell death. In eukaryotes, both the formation and repair of UV damage occur in the context of chromatin, in which genomic DNA is packaged with histones into nucleosomes and higher order chromatin structures. Here, we review how chromatin impacts the formation of UV photoproducts in eukaryotic cells. We describe the initial discovery that nucleosomes and other DNA binding proteins induce characteristic "photofootprints" during the formation of UV photoproducts. We also describe recent progress in genomewide methods for mapping UV damage, which echoes early biochemical studies, and highlights the role of nucleosomes and transcription factors in UV damage formation and repair at unprecedented resolution. Finally, we discuss our current understanding of how the distribution and repair of UV-induced DNA damage influence mutagenesis in human skin cancers.
© 2016 The American Society of Photobiology.
Figures


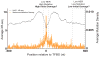
References
-
- van Holde KE. Springer Series in Molecular Biology. Springer; New York,, New York, NY: 1989. Chromatin; p. 1. online resource.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

