Structures and stabilization of kinetoplastid-specific split rRNAs revealed by comparing leishmanial and human ribosomes
- PMID: 27752045
- PMCID: PMC5071889
- DOI: 10.1038/ncomms13223
Structures and stabilization of kinetoplastid-specific split rRNAs revealed by comparing leishmanial and human ribosomes
Abstract
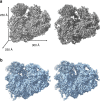
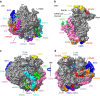
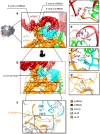
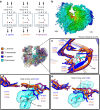
The recent success in ribosome structure determination by cryoEM has opened the door to defining structural differences between ribosomes of pathogenic organisms and humans and to understand ribosome-targeting antibiotics. Here, by direct electron-counting cryoEM, we have determined the structures of the Leishmania donovani and human ribosomes at 2.9 Å and 3.6 Å, respectively. Our structure of the leishmanial ribosome elucidates the organization of the six fragments of its large subunit rRNA (as opposed to a single 28S rRNA in most eukaryotes, including humans) and reveals atomic details of a unique 20 amino acid extension of the uL13 protein that pins down the ends of three of the rRNA fragments. The structure also fashions many large rRNA expansion segments. Direct comparison of our human and leishmanial ribosome structures at the decoding A-site sheds light on how the bacterial ribosome-targeting drug paromomycin selectively inhibits the eukaryotic L. donovani, but not human, ribosome.
Figures



References
-
- World Health Organization. Control of the leishmaniases. World Health Organization Tech Rep Ser 949, 67–71 (2010). - PubMed
-
- Anger A. M. et al. Structures of the human and Drosophila 80S ribosome. Nature 497, 80–85 (2013). - PubMed
-
- Khatter H., Myasnikov A. G., Natchiar S. K. & Klaholz B. P. Structure of the human 80S ribosome. Nature 520, 640–645 (2015). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

