An Intronic Enhancer Element Regulates Angiotensin II Type 2 Receptor Expression during Satellite Cell Differentiation, and Its Activity Is Suppressed in Congestive Heart Failure
- PMID: 27756842
- PMCID: PMC5207256
- DOI: 10.1074/jbc.M116.752501
An Intronic Enhancer Element Regulates Angiotensin II Type 2 Receptor Expression during Satellite Cell Differentiation, and Its Activity Is Suppressed in Congestive Heart Failure
Abstract
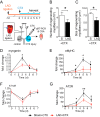
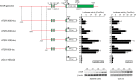
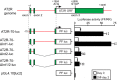
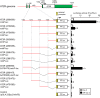
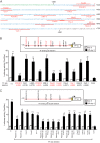
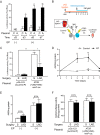
Patients with advanced congestive heart failure (CHF) or chronic kidney disease often have increased angiotensin II (Ang II) levels and cachexia. We previously demonstrated that Ang II, via its type 1 receptor, causes muscle protein breakdown and apoptosis and inhibits satellite cell (SC) proliferation and muscle regeneration, likely contributing to cachexia in CHF and chronic kidney disease. In contrast, Ang II, via its type 2 receptor (AT2R) expression, is robustly induced during SC differentiation, and it potentiates muscle regeneration. To understand the mechanisms regulating AT2R expression and its potential role in muscle regeneration in chronic diseases, we used a mouse model of CHF and found that muscle regeneration was markedly reduced and that this was accompanied by blunted increase of AT2R expression. We performed AT2R promoter reporter analysis during satellite cell differentiation and found that the 70 bp upstream of the AT2R transcription start site contain a core promoter region, and regions upstream of 70 bp to 3 kbp are dispensable for AT2R induction. Instead, AT2R intron 2 acts as a transcriptional enhancer during SC differentiation. Further deletion/mutation analysis revealed that multiple transcription factor binding sites in the +286/+690 region within intron 2 coordinately regulate AT2R transcription. Importantly, +286/+690 enhancer activity was suppressed in CHF mouse skeletal muscle, suggesting that AT2R expression is suppressed in CHF via inhibition of AT2R intronic enhancer activity, leading to lowered muscle regeneration. Thus targeting intron 2 enhancer element could lead to the development of a novel intervention to increase AT2R expression in SCs and potentiate skeletal muscle regenerative capacity in chronic diseases.
Keywords: angiotensin II; heart failure; muscle regeneration; skeletal muscle; stem cells; transcription enhancer.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


References
-
- Anker S. D., Negassa A., Coats A. J., Afzal R., Poole-Wilson P. A., Cohn J. N., and Yusuf S. (2003) Prognostic importance of weight loss in chronic heart failure and the effect of treatment with angiotensin-converting-enzyme inhibitors: an observational study. Lancet 361, 1077–1083 - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

