Central Metabolic Responses to Ozone and Herbivory Affect Photosynthesis and Stomatal Closure
- PMID: 27758847
- PMCID: PMC5100778
- DOI: 10.1104/pp.16.01318
Central Metabolic Responses to Ozone and Herbivory Affect Photosynthesis and Stomatal Closure
Abstract
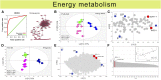
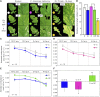
Plants have evolved adaptive mechanisms that allow them to tolerate a continuous range of abiotic and biotic stressors. Tropospheric ozone (O3), a global anthropogenic pollutant, directly affects living organisms and ecosystems, including plant-herbivore interactions. In this study, we investigate the stress responses of Brassica nigra (wild black mustard) exposed consecutively to O3 and the specialist herbivore Pieris brassicae Transcriptomics and metabolomics data were evaluated using multivariate, correlation, and network analyses for the O3 and herbivory responses. O3 stress symptoms resembled those of senescence and phosphate starvation, while a sequential shift from O3 to herbivory induced characteristic plant defense responses, including a decrease in central metabolism, induction of the jasmonic acid/ethylene pathways, and emission of volatiles. Omics network and pathway analyses predicted a link between glycerol and central energy metabolism that influences the osmotic stress response and stomatal closure. Further physiological measurements confirmed that while O3 stress inhibited photosynthesis and carbon assimilation, sequential herbivory counteracted the initial responses induced by O3, resulting in a phenotype similar to that observed after herbivory alone. This study clarifies the consequences of multiple stress interactions on a plant metabolic system and also illustrates how omics data can be integrated to generate new hypotheses in ecology and plant physiology.
© 2016 American Society of Plant Biologists. All Rights Reserved.
Figures







Similar articles
-
The phytotoxic air-pollutant O3 enhances the emission of herbivore-induced volatile organic compounds (VOCs) and affects the susceptibility of black mustard plants to pest attack.Environ Pollut. 2020 Oct;265(Pt A):115030. doi: 10.1016/j.envpol.2020.115030. Epub 2020 Jun 17. Environ Pollut. 2020. PMID: 32806411
-
Water use strategy affects avoidance of ozone stress by stomatal closure in Mediterranean trees-A modelling analysis.Plant Cell Environ. 2020 Mar;43(3):611-623. doi: 10.1111/pce.13700. Epub 2020 Jan 10. Plant Cell Environ. 2020. PMID: 31834637
-
Elevated Ozone Modulates Herbivore-Induced Volatile Emissions of Brassica nigra and Alters a Tritrophic Interaction.J Chem Ecol. 2016 May;42(5):368-81. doi: 10.1007/s10886-016-0697-8. Epub 2016 May 11. J Chem Ecol. 2016. PMID: 27167383
-
Ozone Induced Stomatal Regulations, MAPK and Phytohormone Signaling in Plants.Int J Mol Sci. 2021 Jun 11;22(12):6304. doi: 10.3390/ijms22126304. Int J Mol Sci. 2021. PMID: 34208343 Free PMC article. Review.
-
Stomata-mediated interactions between plants, herbivores, and the environment.Trends Plant Sci. 2022 Mar;27(3):287-300. doi: 10.1016/j.tplants.2021.08.017. Epub 2021 Sep 24. Trends Plant Sci. 2022. PMID: 34580024 Review.
Cited by
-
Combined Acute Ozone and Water Stress Alters the Quantitative Relationships between O3 Uptake, Photosynthetic Characteristics and Volatile Emissions in Brassica nigra.Molecules. 2021 May 23;26(11):3114. doi: 10.3390/molecules26113114. Molecules. 2021. PMID: 34070994 Free PMC article.
-
Single, but not dual, attack by a biotrophic pathogen and a sap-sucking insect affects the oak leaf metabolome.Front Plant Sci. 2022 Aug 3;13:897186. doi: 10.3389/fpls.2022.897186. eCollection 2022. Front Plant Sci. 2022. PMID: 35991442 Free PMC article.
-
Does ozone exposure affect herbivore-induced plant volatile emissions differently in wild and cultivated plants?Environ Sci Pollut Res Int. 2020 Aug;27(24):30448-30459. doi: 10.1007/s11356-020-09320-z. Epub 2020 May 28. Environ Sci Pollut Res Int. 2020. PMID: 32468369 Free PMC article.
-
Terpene Composition Complexity Controls Secondary Organic Aerosol Yields from Scots Pine Volatile Emissions.Sci Rep. 2018 Feb 14;8(1):3053. doi: 10.1038/s41598-018-21045-1. Sci Rep. 2018. PMID: 29445182 Free PMC article.
-
Legacy effects of historical grazing alter leaf stomatal characteristics in progeny plants.PeerJ. 2020 Jun 17;8:e9266. doi: 10.7717/peerj.9266. eCollection 2020. PeerJ. 2020. PMID: 32596041 Free PMC article.
References
-
- Anderson JP, Badruzsaufari E, Schenk PM, Manners JM, Desmond OJ, Ehlert C, Maclean DJ, Ebert PR, Kazan K (2004) Antagonistic interaction between abscisic acid and jasmonate-ethylene signaling pathways modulates defense gene expression and disease resistance in Arabidopsis. Plant Cell 16: 3460–3479 - PMC - PubMed
-
- Ansari M, Lee RH, Chen SC (2005) A novel senescence‐associated gene encoding γ‐aminobutyric acid (GABA):pyruvate transaminase is upregulated during rice leaf senescence. Physiol Plant 123: 1–8
-
- Ashmore MR. (2005) Assessing the future global impacts of ozone on vegetation. Plant Cell Environ 28: 949–964
-
- Assenov Y, Ramírez F, Schelhorn SE, Lengauer T, Albrecht M (2008) Computing topological parameters of biological networks. Bioinformatics 24: 282–284 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

