Mechanism for DNA transposons to generate introns on genomic scales
- PMID: 27760113
- PMCID: PMC5684705
- DOI: 10.1038/nature20110
Mechanism for DNA transposons to generate introns on genomic scales
Abstract
The discovery of introns four decades ago was one of the most unexpected findings in molecular biology. Introns are sequences interrupting genes that must be removed as part of messenger RNA production. Genome sequencing projects have shown that most eukaryotic genes contain at least one intron, and frequently many. Comparison of these genomes reveals a history of long evolutionary periods during which few introns were gained, punctuated by episodes of rapid, extensive gain. However, although several detailed mechanisms for such episodic intron generation have been proposed, none has been empirically supported on a genomic scale. Here we show how short, non-autonomous DNA transposons independently generated hundreds to thousands of introns in the prasinophyte Micromonas pusilla and the pelagophyte Aureococcus anophagefferens. Each transposon carries one splice site. The other splice site is co-opted from the gene sequence that is duplicated upon transposon insertion, allowing perfect splicing out of the RNA. The distributions of sequences that can be co-opted are biased with respect to codons, and phasing of transposon-generated introns is similarly biased. These transposons insert between pre-existing nucleosomes, so that multiple nearby insertions generate nucleosome-sized intervening segments. Thus, transposon insertion and sequence co-option may explain the intron phase biases and prevalence of nucleosome-sized exons observed in eukaryotes. Overall, the two independent examples of proliferating elements illustrate a general DNA transposon mechanism that can plausibly account for episodes of rapid, extensive intron gain during eukaryotic evolution.
Figures




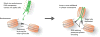







References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

