Transferring entropy to the realm of GxG interactions
- PMID: 27769993
- PMCID: PMC5862307
- DOI: 10.1093/bib/bbw086
Transferring entropy to the realm of GxG interactions
Abstract
Genome-wide association studies are moving to genome-wide interaction studies, as the genetic background of many diseases appears to be more complex than previously supposed. Thus, many statistical approaches have been proposed to detect gene-gene (GxG) interactions, among them numerous information theory-based methods, inspired by the concept of entropy. These are suggested as particularly powerful and, because of their nonlinearity, as better able to capture nonlinear relationships between genetic variants and/or variables. However, the introduced entropy-based estimators differ to a surprising extent in their construction and even with respect to the basic definition of interactions. Also, not every entropy-based measure for interaction is accompanied by a proper statistical test. To shed light on this, a systematic review of the literature is presented answering the following questions: (1) How are GxG interactions defined within the framework of information theory? (2) Which entropy-based test statistics are available? (3) Which underlying distribution do the test statistics follow? (4) What are the given strengths and limitations of these test statistics?
Keywords: entropy; estimation; genetic interactions; information theory.
© The Author 2016. Published by Oxford University Press.
Figures


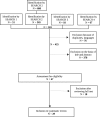
References
-
- Calle ML, Urrea V, Vellalta G, et al.Improving strategies for detecting genetic patterns of disease susceptibility in association studies. Stat Med 2008;27:6532–46. - PubMed
-
- Breiman L. Random forests. Mach Learn 2001;45:5–32.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

