Transcriptome Sequencing Identified Genes and Gene Ontologies Associated with Early Freezing Tolerance in Maize
- PMID: 27774095
- PMCID: PMC5054024
- DOI: 10.3389/fpls.2016.01477
Transcriptome Sequencing Identified Genes and Gene Ontologies Associated with Early Freezing Tolerance in Maize
Abstract
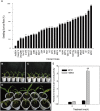
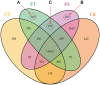
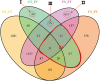
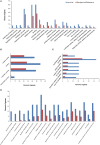
Originating in a tropical climate, maize has faced great challenges as cultivation has expanded to the majority of the world's temperate zones. In these zones, frost and cold temperatures are major factors that prevent maize from reaching its full yield potential. Among 30 elite maize inbred lines adapted to northern China, we identified two lines of extreme, but opposite, freezing tolerance levels-highly tolerant and highly sensitive. During the seedling stage of these two lines, we used RNA-seq to measure changes in maize whole genome transcriptome before and after freezing treatment. In total, 19,794 genes were expressed, of which 4550 exhibited differential expression due to either treatment (before or after freezing) or line type (tolerant or sensitive). Of the 4550 differently expressed genes, 948 exhibited differential expression due to treatment within line or lines under freezing condition. Analysis of gene ontology found that these 948 genes were significantly enriched for binding functions (DNA binding, ATP binding, and metal ion binding), protein kinase activity, and peptidase activity. Based on their enrichment, literature support, and significant levels of differential expression, 30 of these 948 genes were selected for quantitative real-time PCR (qRT-PCR) validation. The validation confirmed our RNA-Seq-based findings, with squared correlation coefficients of 80% and 50% in the tolerance and sensitive lines, respectively. This study provided valuable resources for further studies to enhance understanding of the molecular mechanisms underlying maize early freezing response and enable targeted breeding strategies for developing varieties with superior frost resistance to achieve yield potential.
Keywords: DEGs; RNA-seq; freezing stress; maize; molecular mechanism.
Figures
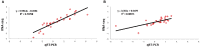
References
-
- Adamczyk J., Królikowski Z. (1998). From open pollinated varieties to single crosses: rapid development in Polish maize breeding, in Crop Development for Cool and Wet European Climate, eds Sowiński P., Zagdańska B., Aniol A., Pithan K. (Brussels: European Cooperation in the Field of Scientific and Technical Research, Commission of European Communities; ), 65–69.
-
- Ascencio-Ibáñez J. T., Sozzani R., Lee T.-J., Chu T.-M., Wolfinger R. D., Cella R., et al. . (2008). Global analysis of Arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection. Plant Physiol. 148, 436–454. 10.1104/pp.108.121038 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

