The effects of sampling strategy on the quality of reconstruction of viral population dynamics using Bayesian skyline family coalescent methods: A simulation study
- PMID: 27774296
- PMCID: PMC4989886
- DOI: 10.1093/ve/vew003
The effects of sampling strategy on the quality of reconstruction of viral population dynamics using Bayesian skyline family coalescent methods: A simulation study
Abstract
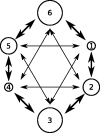
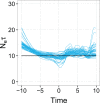
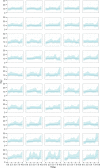
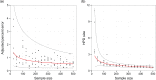
The ongoing large-scale increase in the total amount of genetic data for viruses and other pathogens has led to a situation in which it is often not possible to include every available sequence in a phylogenetic analysis and expect the procedure to complete in reasonable computational time. This raises questions about how a set of sequences should be selected for analysis, particularly if the data are used to infer more than just the phylogenetic tree itself. The design of sampling strategies for molecular epidemiology has been a neglected field of research. This article describes a large-scale simulation exercise that was undertaken to select an appropriate strategy when using the GMRF skygrid, one of the Bayesian skyline family of coalescent methods, in order to reconstruct past population dynamics. The simulated scenarios were intended to represent sampling for the population of an endemic virus across multiple geographical locations. Large phylogenies were simulated under a coalescent or structured coalescent model and sequences simulated from these trees; the resulting datasets were then downsampled for analyses according to a variety of schemes. Variation in results between different replicates of the same scheme was not insignificant, and as a result, we recommend that where possible analyses are repeated with different datasets in order to establish that elements of a reconstruction are not simply the result of the particular set of samples selected. We show that an individual stochastic choice of sequences can introduce spurious behaviour in the median line of the skygrid plot and that even marginal likelihood estimation can suggest complicated dynamics that were not in fact present. We recommend that the median line should not be used to infer historical events on its own. Sampling sequences with uniform probability with respect to both time and spatial location (deme) never performed worse than sampling with probability proportional to the effective population size at that time and in that location and frequently was superior. As a result, we recommend this approach in the design of future studies. We also confirm that the inclusion of many recent sequences from a single geographical location in an analysis tends to result in a spurious bottleneck effect in the reconstruction and caution against interpreting this as genuine.
Keywords: coalescent; phylodynamics; sampling; simulation.
Figures

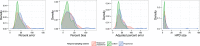

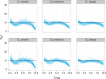
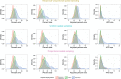
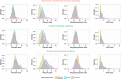

Similar articles
-
Jointly Inferring the Dynamics of Population Size and Sampling Intensity from Molecular Sequences.Mol Biol Evol. 2020 Aug 1;37(8):2414-2429. doi: 10.1093/molbev/msaa016. Mol Biol Evol. 2020. PMID: 32003829 Free PMC article.
-
The Impact of the Tree Prior on Molecular Dating of Data Sets Containing a Mixture of Inter- and Intraspecies Sampling.Syst Biol. 2017 May 1;66(3):413-425. doi: 10.1093/sysbio/syw095. Syst Biol. 2017. PMID: 27798404
-
Hamiltonian Monte Carlo sampling to estimate past population dynamics using the skygrid coalescent model in a Bayesian phylogenetics framework.Wellcome Open Res. 2020 Mar 30;5:53. doi: 10.12688/wellcomeopenres.15770.1. eCollection 2020. Wellcome Open Res. 2020. PMID: 32923688 Free PMC article.
-
Challenges in Species Tree Estimation Under the Multispecies Coalescent Model.Genetics. 2016 Dec;204(4):1353-1368. doi: 10.1534/genetics.116.190173. Genetics. 2016. PMID: 27927902 Free PMC article. Review.
-
Skyline-plot methods for estimating demographic history from nucleotide sequences.Mol Ecol Resour. 2011 May;11(3):423-34. doi: 10.1111/j.1755-0998.2011.02988.x. Epub 2011 Feb 6. Mol Ecol Resour. 2011. PMID: 21481200 Review.
Cited by
-
Accounting for spatial sampling patterns in Bayesian phylogeography.Proc Natl Acad Sci U S A. 2021 Dec 28;118(52):e2105273118. doi: 10.1073/pnas.2105273118. Proc Natl Acad Sci U S A. 2021. PMID: 34930835 Free PMC article.
-
Evidence for a recombinant origin of HIV-1 Group M from genomic variation.Virus Evol. 2019 Jan 22;5(1):vey039. doi: 10.1093/ve/vey039. eCollection 2019 Jan. Virus Evol. 2019. PMID: 30687518 Free PMC article.
-
Origins and Evolution of Seasonal Human Coronaviruses.Viruses. 2022 Jul 15;14(7):1551. doi: 10.3390/v14071551. Viruses. 2022. PMID: 35891531 Free PMC article.
-
Reconstructing the evolutionary history of pandemic foot-and-mouth disease viruses: the impact of recombination within the emerging O/ME-SA/Ind-2001 lineage.Sci Rep. 2018 Oct 2;8(1):14693. doi: 10.1038/s41598-018-32693-8. Sci Rep. 2018. PMID: 30279570 Free PMC article.
-
Genome-wide phylodynamic approach reveals the epidemic dynamics of the main Mycoplasma bovis subtype circulating in France.Microb Genom. 2023 Jul;9(7):mgen001067. doi: 10.1099/mgen.0.001067. Microb Genom. 2023. PMID: 37486749 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

