An Interaction with Ewing's Sarcoma Breakpoint Protein EWS Defines a Specific Oncogenic Mechanism of ETS Factors Rearranged in Prostate Cancer
- PMID: 27783944
- PMCID: PMC5123826
- DOI: 10.1016/j.celrep.2016.10.001
An Interaction with Ewing's Sarcoma Breakpoint Protein EWS Defines a Specific Oncogenic Mechanism of ETS Factors Rearranged in Prostate Cancer
Abstract
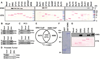
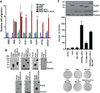
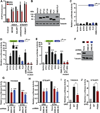
More than 50% of prostate tumors have a chromosomal rearrangement resulting in aberrant expression of an oncogenic ETS family transcription factor. However, mechanisms that differentiate the function of oncogenic ETS factors expressed in prostate tumors from non-oncogenic ETS factors expressed in normal prostate are unknown. Here, we find that four oncogenic ETS (ERG, ETV1, ETV4, and ETV5), and no other ETS, interact with the Ewing's sarcoma breakpoint protein, EWS. This EWS interaction was necessary and sufficient for oncogenic ETS functions including gene activation, cell migration, clonogenic survival, and transformation. Significantly, the EWS interacting region of ERG has no homology with that of ETV1, ETV4, and ETV5. Therefore, this finding may explain how divergent ETS factors have a common oncogenic function. Strikingly, EWS is fused to various ETS factors by the chromosome translocations that cause Ewing's sarcoma. Therefore, these findings link oncogenic ETS function in both prostate cancer and Ewing's sarcoma.
Keywords: ERG; ETS; EWS; Ewing’s sarcoma; prostate cancer.
Copyright © 2016 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures



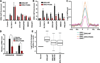
Similar articles
-
Expression of EWS-ETS fusions in NIH3T3 cells reveals significant differences to Ewing's sarcoma.Cell Cycle. 2006 Dec;5(23):2753-9. doi: 10.4161/cc.5.23.3505. Epub 2006 Dec 1. Cell Cycle. 2006. PMID: 17172842
-
Promiscuous partnerships in Ewing's sarcoma.Cancer Genet. 2011 Jul;204(7):351-65. doi: 10.1016/j.cancergen.2011.07.008. Cancer Genet. 2011. PMID: 21872822 Free PMC article. Review.
-
Cooperative DNA binding with AP-1 proteins is required for transformation by EWS-Ets fusion proteins.Mol Cell Biol. 2006 Apr;26(7):2467-78. doi: 10.1128/MCB.26.7.2467-2478.2006. Mol Cell Biol. 2006. PMID: 16537893 Free PMC article.
-
Caveolin-1 (CAV1) is a target of EWS/FLI-1 and a key determinant of the oncogenic phenotype and tumorigenicity of Ewing's sarcoma cells.Cancer Res. 2006 Oct 15;66(20):9937-47. doi: 10.1158/0008-5472.CAN-06-0927. Cancer Res. 2006. PMID: 17047056
-
Context matters: the hen or egg problem in Ewing's sarcoma.Semin Cancer Biol. 2005 Jun;15(3):189-96. doi: 10.1016/j.semcancer.2005.01.004. Semin Cancer Biol. 2005. PMID: 15826833 Review.
Cited by
-
Common ELF1 deletion in prostate cancer bolsters oncogenic ETS function, inhibits senescence and promotes docetaxel resistance.Genes Cancer. 2018 May;9(5-6):198-214. doi: 10.18632/genesandcancer.182. Genes Cancer. 2018. PMID: 30603056 Free PMC article.
-
Ras and Wnt Interaction Contribute in Prostate Cancer Bone Metastasis.Molecules. 2020 May 20;25(10):2380. doi: 10.3390/molecules25102380. Molecules. 2020. PMID: 32443915 Free PMC article. Review.
-
Novel potent anti-STEAP1 bispecific antibody to redirect T cells for cancer immunotherapy.J Immunother Cancer. 2021 Sep;9(9):e003114. doi: 10.1136/jitc-2021-003114. J Immunother Cancer. 2021. PMID: 34497115 Free PMC article.
-
Past, Current, and Future Strategies to Target ERG Fusion-Positive Prostate Cancer.Cancers (Basel). 2022 Feb 22;14(5):1118. doi: 10.3390/cancers14051118. Cancers (Basel). 2022. PMID: 35267426 Free PMC article. Review.
-
Discovery of key genes as novel biomarkers specifically associated with HPV-negative cervical cancer.Mol Ther Methods Clin Dev. 2021 Apr 6;21:492-506. doi: 10.1016/j.omtm.2021.03.026. eCollection 2021 Jun 11. Mol Ther Methods Clin Dev. 2021. PMID: 33997099 Free PMC article.
References
-
- Araya N, Hirota K, Shimamoto Y, Miyagishi M, Yoshida E, Ishida J, Kaneko S, Kaneko M, Nakajima T, Fukamizu A. Cooperative interaction of EWS with CREB-binding protein selectively activates hepatocyte nuclear factor 4-mediated transcription. J. Biol. Chem. 2003;278:5427–5432. - PubMed
-
- Arvand A, Denny CT. Biology of EWS/ETS fusions in Ewing’s family tumors. Oncogene. 2001;20:5747–5754. - PubMed
-
- Bertolotti A, Melot T, Acker J, Vigneron M, Delattre O, Tora L. EWS, but not EWS-FLI-1, is associated with both TFIID and RNA polymerase II: interactions between two members of the TET family, EWS and hTAFII68, and subunits of TFIID and RNA polymerase II complexes. Mol. Cell. Biol. 1998;18:1489–1497. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

