Mitochondrial function controls intestinal epithelial stemness and proliferation
- PMID: 27786175
- PMCID: PMC5080445
- DOI: 10.1038/ncomms13171
Mitochondrial function controls intestinal epithelial stemness and proliferation
Abstract
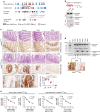
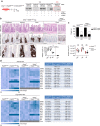
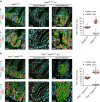
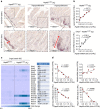
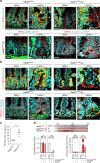
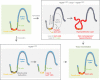
Control of intestinal epithelial stemness is crucial for tissue homeostasis. Disturbances in epithelial function are implicated in inflammatory and neoplastic diseases of the gastrointestinal tract. Here we report that mitochondrial function plays a critical role in maintaining intestinal stemness and homeostasis. Using intestinal epithelial cell (IEC)-specific mouse models, we show that loss of HSP60, a mitochondrial chaperone, activates the mitochondrial unfolded protein response (MT-UPR) and results in mitochondrial dysfunction. HSP60-deficient crypts display loss of stemness and cell proliferation, accompanied by epithelial release of WNT10A and RSPO1. Sporadic failure of Cre-mediated Hsp60 deletion gives rise to hyperproliferative crypt foci originating from OLFM4+ stem cells. These effects are independent of the MT-UPR-associated transcription factor CHOP. In conclusion, compensatory hyperproliferation of HSP60+ escaper stem cells suggests paracrine release of WNT-related factors from HSP60-deficient, functionally impaired IEC to be pivotal in the control of the proliferative capacity of the stem cell niche.
Figures




References
-
- Sartor R. B. Mechanisms of disease: pathogenesis of Crohn's disease and ulcerative colitis. Nat. Clin. Pract. Gastroenterol. Hepatol. 3, 390–407 (2006). - PubMed
-
- Henderson P., van Limbergen J. E., Schwarze J. & Wilson D. C. Function of the intestinal epithelium and its dysregulation in inflammatory bowel disease. Inflamm. Bowel Dis. 17, 382–395 (2011). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

