AP1 transcription factors are required to maintain the peripheral taste system
- PMID: 27787515
- PMCID: PMC5133999
- DOI: 10.1038/cddis.2016.343
AP1 transcription factors are required to maintain the peripheral taste system
Abstract
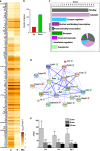
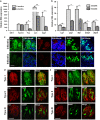
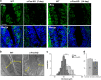
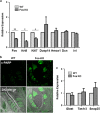
The sense of taste is used by organisms to achieve the optimal nutritional requirement and avoid potentially toxic compounds. In the oral cavity, taste receptor cells are grouped together in taste buds that are present in specialized taste papillae in the tongue. Taste receptor cells are the cells that detect chemicals in potential food items and transmit that information to gustatory nerves that convey the taste information to the brain. As taste cells are in contact with the external environment, they can be damaged and are routinely replaced throughout an organism's lifetime to maintain functionality. However, this taste cell turnover loses efficiency over time resulting in a reduction in taste ability. Currently, very little is known about the mechanisms that regulate the renewal and maintenance of taste cells. We therefore performed RNA-sequencing analysis on isolated taste cells from 2 and 6-month-old mice to determine how alterations in the taste cell-transcriptome regulate taste cell maintenance and function in adults. We found that the activator protein-1 (AP1) transcription factors (c-Fos, Fosb and c-Jun) and genes associated with this pathway were significantly downregulated in taste cells by 6 months and further declined at 12 months. We generated conditional c-Fos-knockout mice to target K14-expressing cells, including differentiating taste cells. c-Fos deletion caused a severe perturbation in taste bud structure and resulted in a significant reduction in the taste bud size. c-Fos deletion also affected taste cell turnover as evident by a decrease in proliferative marker, and upregulation of the apoptotic marker cleaved-PARP. Thus, AP1 factors are important regulators of adult taste cell renewal and their downregulation negatively impacts taste maintenance.
Figures

Similar articles
-
Expression of Sox2 in mouse taste buds and its relation to innervation.Cell Tissue Res. 2008 Jun;332(3):393-401. doi: 10.1007/s00441-008-0600-1. Epub 2008 Apr 1. Cell Tissue Res. 2008. PMID: 18379823
-
Expression of glial cell line-derived neurotrophic factor (GDNF) and GDNF family receptor alpha1 in mouse taste bud cells after denervation.Anat Sci Int. 2005 Jun;80(2):105-10. doi: 10.1111/j.1447-073x.2005.00105.x. Anat Sci Int. 2005. PMID: 15960316
-
Mash1 is required for the differentiation of AADC-positive type III cells in mouse taste buds.Dev Dyn. 2011 Apr;240(4):775-84. doi: 10.1002/dvdy.22576. Epub 2011 Feb 14. Dev Dyn. 2011. PMID: 21322090
-
Progress and renewal in gustation: new insights into taste bud development.Development. 2015 Nov 1;142(21):3620-9. doi: 10.1242/dev.120394. Development. 2015. PMID: 26534983 Free PMC article. Review.
-
Developing and regenerating a sense of taste.Curr Top Dev Biol. 2015;111:401-19. doi: 10.1016/bs.ctdb.2014.11.012. Epub 2015 Jan 20. Curr Top Dev Biol. 2015. PMID: 25662267 Free PMC article. Review.
Cited by
-
A subset of broadly responsive Type III taste cells contribute to the detection of bitter, sweet and umami stimuli.PLoS Genet. 2020 Aug 13;16(8):e1008925. doi: 10.1371/journal.pgen.1008925. eCollection 2020 Aug. PLoS Genet. 2020. PMID: 32790785 Free PMC article.
-
Nutrient Sensing by Lingual G-Protein-Coupled Taste Receptors.Methods Mol Biol. 2025;2882:317-327. doi: 10.1007/978-1-0716-4284-9_16. Methods Mol Biol. 2025. PMID: 39992517
-
c-Fos downregulation positively regulates EphA5 expression in a congenital hypothyroidism rat model.J Mol Histol. 2018 Apr;49(2):147-155. doi: 10.1007/s10735-018-9754-7. Epub 2018 Jan 12. J Mol Histol. 2018. PMID: 29330744
-
TRPM4 and TRPM5 are both required for normal signaling in taste receptor cells.Proc Natl Acad Sci U S A. 2018 Jan 23;115(4):E772-E781. doi: 10.1073/pnas.1718802115. Epub 2018 Jan 8. Proc Natl Acad Sci U S A. 2018. PMID: 29311301 Free PMC article.
-
The stability of tastant detection by mouse lingual chemosensory tissue requires Regulator of G protein Signaling-21 (RGS21).Chem Senses. 2021 Jan 1;46:bjab048. doi: 10.1093/chemse/bjab048. Chem Senses. 2021. PMID: 34718440 Free PMC article.
References
-
- Grando LJ, Mello AL, Salvato L, Brancher AP, Del Moral JA, Steffenello-Durigon G. Impact of leukemia and lymphoma chemotherapy on oral cavity and quality of life. Spec Care Dentist 2015. (doi:10.1111/scd.12113). - PubMed
-
- Roura E, Foster S, Winklebach A, Navarro M, Thomas W, Campbell K et al. Taste and hypertension in humans: targeting cardiovascular disease. Curr Pharm Des 2016; 22: 2290–2305. - PubMed
-
- Mukherjee N, Delay ER. Cyclophosphamide-induced disruption of umami taste functions and taste epithelium. Neuroscience 2011; 192: 732–745. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

