Gene transfers from diverse bacteria compensate for reductive genome evolution in the chromatophore of Paulinella chromatophora
- PMID: 27791007
- PMCID: PMC5087059
- DOI: 10.1073/pnas.1608016113
Gene transfers from diverse bacteria compensate for reductive genome evolution in the chromatophore of Paulinella chromatophora
Abstract
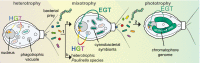
Plastids, the photosynthetic organelles, originated >1 billion y ago via the endosymbiosis of a cyanobacterium. The resulting proliferation of primary producers fundamentally changed global ecology. Endosymbiotic gene transfer (EGT) from the intracellular cyanobacterium to the nucleus is widely recognized as a critical factor in the evolution of photosynthetic eukaryotes. The contribution of horizontal gene transfers (HGTs) from other bacteria to plastid establishment remains more controversial. A novel perspective on this issue is provided by the amoeba Paulinella chromatophora, which contains photosynthetic organelles (chromatophores) that are only 60-200 million years old. Chromatophore genome reduction entailed the loss of many biosynthetic pathways including those for numerous amino acids and cofactors. How the host cell compensates for these losses remains unknown, because the presence of bacteria in all available P. chromatophora cultures excluded elucidation of the full metabolic capacity and occurrence of HGT in this species. Here we generated a high-quality transcriptome and draft genome assembly from the first bacteria-free P. chromatophora culture to deduce rules that govern organelle integration into cellular metabolism. Our analyses revealed that nuclear and chromatophore gene inventories provide highly complementary functions. At least 229 nuclear genes were acquired via HGT from various bacteria, of which only 25% putatively arose through EGT from the chromatophore genome. Many HGT-derived bacterial genes encode proteins that fill gaps in critical chromatophore pathways/processes. Our results demonstrate a dominant role for HGT in compensating for organelle genome reduction and suggest that phagotrophy may be a major driver of HGT.
Keywords: coevolution; endosymbiosis; genome evolution; horizontal gene transfer; organellogenesis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures







Similar articles
-
Impact of light intensity and quality on chromatophore and nuclear gene expression in Paulinella chromatophora, an amoeba with nascent photosynthetic organelles.Plant J. 2017 Apr;90(2):221-234. doi: 10.1111/tpj.13488. Epub 2017 Mar 22. Plant J. 2017. PMID: 28182317
-
Endosymbiotic gene transfer and transcriptional regulation of transferred genes in Paulinella chromatophora.Mol Biol Evol. 2011 Jan;28(1):407-22. doi: 10.1093/molbev/msq209. Epub 2010 Aug 11. Mol Biol Evol. 2011. PMID: 20702568
-
Massive Protein Import into the Early-Evolutionary-Stage Photosynthetic Organelle of the Amoeba Paulinella chromatophora.Curr Biol. 2017 Sep 25;27(18):2763-2773.e5. doi: 10.1016/j.cub.2017.08.010. Epub 2017 Sep 7. Curr Biol. 2017. PMID: 28889978
-
Paulinella, a model for understanding plastid primary endosymbiosis.J Phycol. 2020 Aug;56(4):837-843. doi: 10.1111/jpy.13003. Epub 2020 May 5. J Phycol. 2020. PMID: 32289879 Free PMC article. Review.
-
Role of horizontal gene transfer in the evolution of photosynthetic eukaryotes and their plastids.Methods Mol Biol. 2009;532:501-15. doi: 10.1007/978-1-60327-853-9_29. Methods Mol Biol. 2009. PMID: 19271204 Review.
Cited by
-
Evolutionary dynamics of the chromatophore genome in three photosynthetic Paulinella species.Sci Rep. 2019 Feb 22;9(1):2560. doi: 10.1038/s41598-019-38621-8. Sci Rep. 2019. PMID: 30796245 Free PMC article.
-
Functional horizontal gene transfer from bacteria to eukaryotes.Nat Rev Microbiol. 2018 Feb;16(2):67-79. doi: 10.1038/nrmicro.2017.137. Epub 2017 Nov 27. Nat Rev Microbiol. 2018. PMID: 29176581 Review.
-
Amoeba Genome Reveals Dominant Host Contribution to Plastid Endosymbiosis.Mol Biol Evol. 2021 Jan 23;38(2):344-357. doi: 10.1093/molbev/msaa206. Mol Biol Evol. 2021. PMID: 32790833 Free PMC article.
-
Pyropia yezoensis genome reveals diverse mechanisms of carbon acquisition in the intertidal environment.Nat Commun. 2020 Aug 12;11(1):4028. doi: 10.1038/s41467-020-17689-1. Nat Commun. 2020. PMID: 32788591 Free PMC article.
-
Are Cyanobacteria an Ancestor of Chloroplasts or Just One of the Gene Donors for Plants and Algae?Genes (Basel). 2021 May 27;12(6):823. doi: 10.3390/genes12060823. Genes (Basel). 2021. PMID: 34071987 Free PMC article.
References
-
- Falkowski PG, et al. The evolution of modern eukaryotic phytoplankton. Science. 2004;305(5682):354–360. - PubMed
-
- Yoon HS, Hackett JD, Ciniglia C, Pinto G, Bhattacharya D. A molecular timeline for the origin of photosynthetic eukaryotes. Mol Biol Evol. 2004;21(5):809–818. - PubMed
-
- Gould SB, Waller RF, McFadden GI. Plastid evolution. Annu Rev Plant Biol. 2008;59:491–517. - PubMed
-
- Schleiff E, Becker T. Common ground for protein translocation: Access control for mitochondria and chloroplasts. Nat Rev Mol Cell Biol. 2011;12(1):48–59. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

