GSDMB induces an asthma phenotype characterized by increased airway responsiveness and remodeling without lung inflammation
- PMID: 27799535
- PMCID: PMC5135378
- DOI: 10.1073/pnas.1610433113
GSDMB induces an asthma phenotype characterized by increased airway responsiveness and remodeling without lung inflammation
Abstract
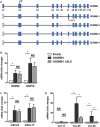
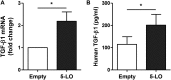
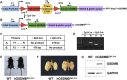
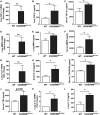
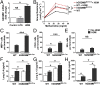
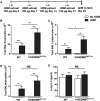
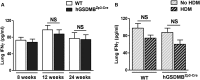
Gasdermin B (GSDMB) on chromosome 17q21 demonstrates a strong genetic linkage to asthma, but its function in asthma is unknown. Here we identified that GSDMB is highly expressed in lung bronchial epithelium in human asthma. Overexpression of GSDMB in primary human bronchial epithelium increased expression of genes important to both airway remodeling [TGF-β1, 5-lipoxygenase (5-LO)] and airway-hyperresponsiveness (AHR) (5-LO). Interestingly, hGSDMBZp3-Cre mice expressing increased levels of the human GSDMB transgene showed a significant spontaneous increase in AHR and a significant spontaneous increase in airway remodeling, with increased smooth muscle mass and increased fibrosis in the absence of airway inflammation. In addition, hGSDMBZp3-Cre mice showed increases in the same remodeling and AHR mediators (TGF-β1, 5-LO) observed in vitro in GSDMB-overexpressing epithelial cells. GSDMB induces TGF-β1 expression via induction of 5-LO, because knockdown of 5-LO in epithelial cells overexpressing GSDMB inhibited TGF-β1 expression. These studies demonstrate that GSDMB, a gene highly linked to asthma but whose function in asthma is previously unknown, regulates AHR and airway remodeling without airway inflammation through a previously unrecognized pathway in which GSDMB induces 5-LO to induce TGF-β1 in bronchial epithelium.
Keywords: GSDMB; airway-hyperresponsiveness; asthma; inflammation; remodeling.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
-
- Moffatt MF, et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature. 2007;448(7152):470–473. - PubMed
-
- Marsh DG, et al. Linkage analysis of IL4 and other chromosome 5q31.1 markers and total serum immunoglobulin E concentrations. Science. 1994;264(5162):1152–1156. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

