RGAugury: a pipeline for genome-wide prediction of resistance gene analogs (RGAs) in plants
- PMID: 27806688
- PMCID: PMC5093994
- DOI: 10.1186/s12864-016-3197-x
RGAugury: a pipeline for genome-wide prediction of resistance gene analogs (RGAs) in plants
Abstract
Background: Resistance gene analogs (RGAs), such as NBS-encoding proteins, receptor-like protein kinases (RLKs) and receptor-like proteins (RLPs), are potential R-genes that contain specific conserved domains and motifs. Thus, RGAs can be predicted based on their conserved structural features using bioinformatics tools. Computer programs have been developed for the identification of individual domains and motifs from the protein sequences of RGAs but none offer a systematic assessment of the different types of RGAs. A user-friendly and efficient pipeline is needed for large-scale genome-wide RGA predictions of the growing number of sequenced plant genomes.
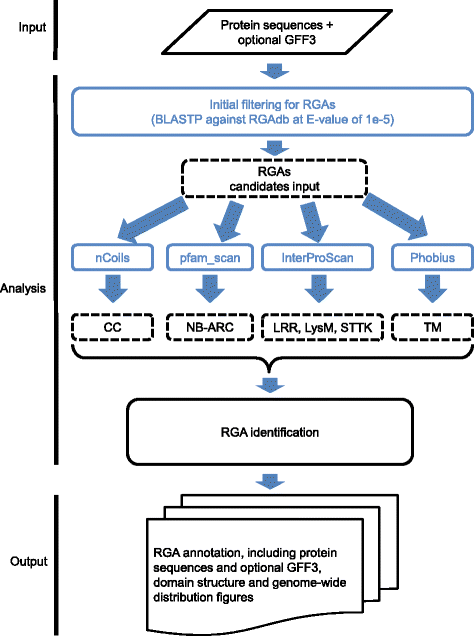
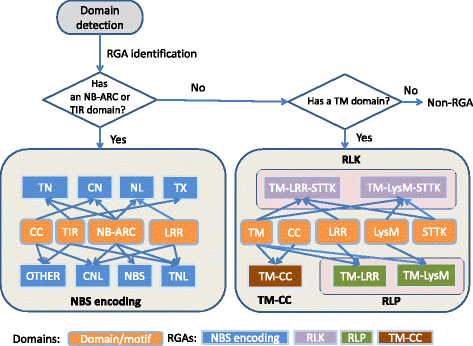
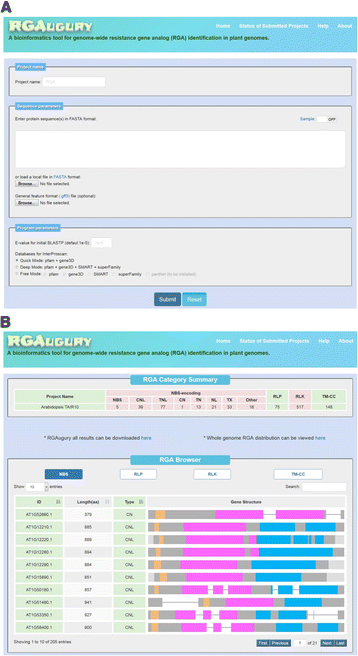
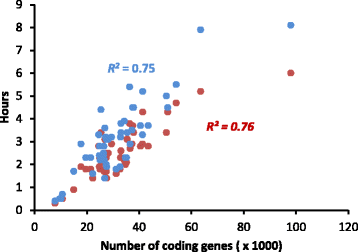
Results: An integrative pipeline, named RGAugury, was developed to automate RGA prediction. The pipeline first identifies RGA-related protein domains and motifs, namely nucleotide binding site (NB-ARC), leucine rich repeat (LRR), transmembrane (TM), serine/threonine and tyrosine kinase (STTK), lysin motif (LysM), coiled-coil (CC) and Toll/Interleukin-1 receptor (TIR). RGA candidates are identified and classified into four major families based on the presence of combinations of these RGA domains and motifs: NBS-encoding, TM-CC, and membrane associated RLP and RLK. All time-consuming analyses of the pipeline are paralleled to improve performance. The pipeline was evaluated using the well-annotated Arabidopsis genome. A total of 98.5, 85.2, and 100 % of the reported NBS-encoding genes, membrane associated RLPs and RLKs were validated, respectively. The pipeline was also successfully applied to predict RGAs for 50 sequenced plant genomes. A user-friendly web interface was implemented to ease command line operations, facilitate visualization and simplify result management for multiple datasets.
Conclusions: RGAugury is an efficiently integrative bioinformatics tool for large scale genome-wide identification of RGAs. It is freely available at Bitbucket: https://bitbucket.org/yaanlpc/rgaugury .
Keywords: Genome-wide prediction; Nucleotide binding site (NBS); Pipeline; Receptor like kinase (RLK); Receptor like protein (RLP); Resistance gene analog (RGA).
Figures
Similar articles
-
Genome survey of resistance gene analogs in sugarcane: genomic features and differential expression of the innate immune system from a smut-resistant genotype.BMC Genomics. 2019 Nov 6;20(1):809. doi: 10.1186/s12864-019-6207-y. BMC Genomics. 2019. PMID: 31694536 Free PMC article.
-
Disease Resistance Gene Analogs (RGAs) in Plants.Int J Mol Sci. 2015 Aug 14;16(8):19248-90. doi: 10.3390/ijms160819248. Int J Mol Sci. 2015. PMID: 26287177 Free PMC article. Review.
-
Exploring Cloned Disease Resistance Gene Homologues and Resistance Gene Analogues in Brassica nigra, Sinapis arvensis, and Sinapis alba: Identification, Characterisation, Distribution, and Evolution.Genes (Basel). 2025 Jul 22;16(8):849. doi: 10.3390/genes16080849. Genes (Basel). 2025. PMID: 40869898
-
Genome-Wide Mining of Disease Resistance Gene Analogs Using Conserved Domains.Methods Mol Biol. 2020;2107:365-375. doi: 10.1007/978-1-0716-0235-5_20. Methods Mol Biol. 2020. PMID: 31893459
-
[Plant resistance genes: molecular and genetic organization, function and evolution].Zh Obshch Biol. 2003 May-Jun;64(3):195-214. Zh Obshch Biol. 2003. PMID: 12815938 Review. Russian.
Cited by
-
Variation in abundance of predicted resistance genes in the Brassica oleracea pangenome.Plant Biotechnol J. 2019 Apr;17(4):789-800. doi: 10.1111/pbi.13015. Epub 2018 May 31. Plant Biotechnol J. 2019. PMID: 30230187 Free PMC article.
-
Genome survey of resistance gene analogs in sugarcane: genomic features and differential expression of the innate immune system from a smut-resistant genotype.BMC Genomics. 2019 Nov 6;20(1):809. doi: 10.1186/s12864-019-6207-y. BMC Genomics. 2019. PMID: 31694536 Free PMC article.
-
Insights into the Genetic Architecture and Genomic Prediction of Powdery Mildew Resistance in Flax (Linum usitatissimum L.).Int J Mol Sci. 2022 Apr 29;23(9):4960. doi: 10.3390/ijms23094960. Int J Mol Sci. 2022. PMID: 35563347 Free PMC article.
-
Identification of powdery mildew resistance QTL in strawberry (Fragaria × ananassa).Theor Appl Genet. 2018 Sep;131(9):1995-2007. doi: 10.1007/s00122-018-3128-0. Epub 2018 Jul 3. Theor Appl Genet. 2018. PMID: 29971472 Free PMC article.
-
Genome-Wide Identification and Evolutionary Analysis of NBS-LRR Genes From Dioscorea rotundata.Front Genet. 2020 May 7;11:484. doi: 10.3389/fgene.2020.00484. eCollection 2020. Front Genet. 2020. PMID: 32457809 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

