Influenza Virus Overcomes Cellular Blocks To Productively Replicate, Impacting Macrophage Function
- PMID: 27807237
- PMCID: PMC5215328
- DOI: 10.1128/JVI.01417-16
Influenza Virus Overcomes Cellular Blocks To Productively Replicate, Impacting Macrophage Function
Abstract
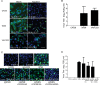
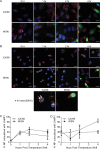
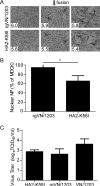
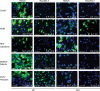
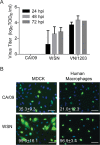
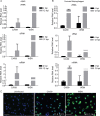
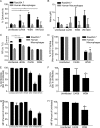
Whether influenza virus replication in macrophages is productive or abortive has been a topic of debate. Utilizing a panel of 28 distinct human, avian, and swine influenza viruses, we found that only a small subset can overcome cellular blocks to productively replicate in murine and primary human macrophages. Murine macrophages have two cellular blocks. The first block is during viral entry, where virions with relatively acid-stable hemagglutinin (HA) proteins are rendered incapable of pH-induced triggering for membrane fusion, resulting in lysosomal degradation. The second block is downstream of viral replication but upstream of late protein synthesis. In contrast, primary human macrophages only have one cellular block that occurs after late protein synthesis. To determine the impact of abortive replication at different stages of the viral life cycle or productive replication on macrophage function, we assessed cytotoxicity, nitric oxide or reactive oxygen species production, and phagocytosis. Intriguingly, productive viral replication decreased phagocytosis of IgG-opsonized bioparticles and Fc receptor CD16 and CD32 surface levels, a function, to our knowledge, never before reported for an RNA virus. These data suggest that replication in macrophages affects cellular function and plays an important role in pathogenesis during infection in vivo IMPORTANCE: Macrophages are a critical first line of defense against respiratory pathogens. Thus, understanding how viruses evade or exploit macrophage function will provide greater insight into viral pathogenicity and antiviral responses. We previously showed that only a subset of highly pathogenic avian (HPAI) H5N1 influenza virus strains could productively replicate in murine macrophages through a hemagglutinin (HA)-mediated mechanism. These studies expand upon this work and demonstrate that productive replication is not specific to unique HPAI H5N1 viruses; an H1N1 strain (A/WSN/33) can also replicate in macrophages. Importantly, we identify two cellular blocks limiting replication that can be overcome by an avian-like pH of activation for nuclear entry and a yet-to-be-identified mechanism(s) to overcome a postnuclear entry block. Overcoming these blocks reduces the cell's ability to phagocytose IgG-opsonized bioparticles by decreasing Fc receptor surface levels, a mechanism previously thought to occur during bacterial and DNA viral infections.
Keywords: influenza; macrophage; replication.
Copyright © 2017 American Society for Microbiology.
Figures


References
-
- Schneider C, Nobs SP, Heer AK, Kurrer M, Klinke G, van Rooijen N, Vogel J, Kopf M. 2014. Alveolar macrophages are essential for protection from respiratory failure and associated morbidity following influenza virus infection. PLoS Pathog 10:e1004053. doi:10.1371/journal.ppat.1004053. - DOI - PMC - PubMed
-
- Cheung CY, Poon LL, Lau AS, Luk W, Lau YL, Shortridge KF, Gordon S, Guan Y, Peiris JS. 2002. Induction of proinflammatory cytokines in human macrophages by influenza A (H5N1) viruses: a mechanism for the unusual severity of human disease? Lancet 360:1831–1837. doi:10.1016/S0140-6736(02)11772-7. - DOI - PubMed
-
- Kobasa D, Jones SM, Shinya K, Kash JC, Copps J, Ebihara H, Hatta Y, Kim JH, Halfmann P, Hatta M, Feldmann F, Alimonti JB, Fernando L, Li Y, Katze MG, Feldmann H, Kawaoka Y. 2007. Aberrant innate immune response in lethal infection of macaques with the 1918 influenza virus. Nature 445:319–323. doi:10.1038/nature05495. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

