Autophagy flux in CA1 neurons of Alzheimer hippocampus: Increased induction overburdens failing lysosomes to propel neuritic dystrophy
- PMID: 27813694
- PMCID: PMC5173282
- DOI: 10.1080/15548627.2016.1239003
Autophagy flux in CA1 neurons of Alzheimer hippocampus: Increased induction overburdens failing lysosomes to propel neuritic dystrophy
Abstract
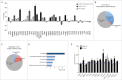
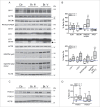
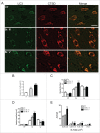
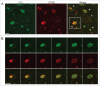
Defective autophagy contributes to Alzheimer disease (AD) pathogenesis although evidence is conflicting on whether multiple stages are impaired. Here, for the first time, we have comprehensively evaluated the entire autophagic process specifically in CA1 pyramidal neurons of hippocampus from early and late-stage AD subjects and nondemented controls. CA1 neurons aspirated by laser capture microdissection were analyzed using a custom-designed microarray comprising 578 neuropathology- and neuroscience-associated genes. Striking upregulation of autophagy-related genes, exceeding that of other gene ontology groups, reflected increases in autophagosome formation and lysosomal biogenesis beginning at early AD stages. Upregulated autophagosome formation was further indicated by elevated gene and protein expression levels for autophagosome components and increased LC3-positive puncta. Increased lysosomal biogenesis was evidenced by activation of MiTF/TFE family transcriptional regulators, particularly TFE3 (transcription factor binding to IGHM enhancer 3) and by elevated expression of their target genes and encoded proteins. Notably, TFEB (transcription factor EB) activation was associated more strongly with glia than neurons. These findings establish that autophagic sequestration is both competent and upregulated in AD. Autophagosome-lysosome fusion is not evidently altered. Despite this early disease response, however, autophagy flux is progressively impeded due to deficient substrate clearance, as reflected by autolysosomal accumulation of LC3-II and SQSTM1/p62 and expansion of autolysosomal size and total area. We propose that sustained induction of autophagy in the face of progressively declining lysosomal clearance of substrates explains the uncommonly robust autophagic pathology and neuritic dystrophy implicated in AD pathogenesis.
Keywords: Alzheimer disease; CA1 pyramidal neurons; MTOR; TFE3; TFEB; autophagosomes; autophagy; dystrophic neurites; hippocampus; lysosomes.
Figures



References
-
- Nixon RA. The role of autophagy in neurodegenerative disease. Nat Med 2013; 19:983-97; PMID:23921753; http://dx.doi.org/10.1038/nm.3232 - DOI - PubMed
-
- Boya P. Lysosomal function and dysfunction: mechanism and disease. Antioxid Redox Signal 2012; 17:766-74; PMID:22098160; http://dx.doi.org/10.1089/ars.2011.4405 - DOI - PubMed
-
- Menzies FM, Moreau K, Rubinsztein DC. Protein misfolding disorders and macroautophagy. Curr Opin Cell Biol 2011; 23:190-7; PMID:21087849; http://dx.doi.org/10.1016/j.ceb.2010.10.010 - DOI - PMC - PubMed
-
- Simonsen A, Cumming RC, Brech A, Isakson P, Schubert DR, Finley KD. Promoting basal levels of autophagy in the nervous system enhances longevity and oxidant resistance in adult Drosophila. Autophagy 2008; 4:176-84; PMID:18059160; http://dx.doi.org/10.4161/auto.5269 - DOI - PubMed
-
- Madeo F, Tavernarakis N, Kroemer G. Can autophagy promote longevity? Nat Cell Biol 2010; 12:842-6; PMID:20811357; http://dx.doi.org/10.1038/ncb0910-842 - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
