Guanine nucleotide exchange factor RABGEF1 regulates keratinocyte-intrinsic signaling to maintain skin homeostasis
- PMID: 27820702
- PMCID: PMC5127679
- DOI: 10.1172/JCI86359
Guanine nucleotide exchange factor RABGEF1 regulates keratinocyte-intrinsic signaling to maintain skin homeostasis
Abstract
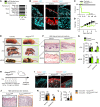
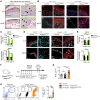
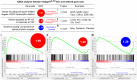
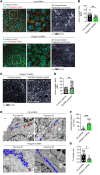
Epidermal keratinocytes form a structural and immune barrier that is essential for skin homeostasis. However, the mechanisms that regulate epidermal barrier function are incompletely understood. Here we have found that keratinocyte-specific deletion of the gene encoding RAB guanine nucleotide exchange factor 1 (RABGEF1, also known as RABEX-5) severely impairs epidermal barrier function in mice and induces an allergic cutaneous and systemic phenotype. RABGEF1-deficient keratinocytes exhibited aberrant activation of the intrinsic IL-1R/MYD88/NF-κB signaling pathway and MYD88-dependent abnormalities in expression of structural proteins that contribute to skin barrier function. Moreover, ablation of MYD88 signaling in RABGEF1-deficient keratinocytes or deletion of Il1r1 restored skin homeostasis and prevented development of skin inflammation. We further demonstrated that epidermal RABGEF1 expression is reduced in skin lesions of humans diagnosed with either atopic dermatitis or allergic contact dermatitis as well as in an inducible mouse model of allergic dermatitis. Our findings reveal a key role for RABGEF1 in dampening keratinocyte-intrinsic MYD88 signaling and sustaining epidermal barrier function in mice, and suggest that dysregulation of RABGEF1 expression may contribute to epidermal barrier dysfunction in allergic skin disorders in mice and humans. Thus, RABGEF1-mediated regulation of IL-1R/MYD88 signaling might represent a potential therapeutic target.
Conflict of interest statement
The authors have declared that no conflict of interest exists.
Figures


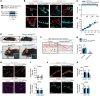



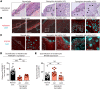

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

