Systematic approaches to identify E3 ligase substrates
- PMID: 27834739
- PMCID: PMC5103871
- DOI: 10.1042/BCJ20160719
Systematic approaches to identify E3 ligase substrates
Abstract
Protein ubiquitylation is a widespread post-translational modification, regulating cellular signalling with many outcomes, such as protein degradation, endocytosis, cell cycle progression, DNA repair and transcription. E3 ligases are a critical component of the ubiquitin proteasome system (UPS), determining the substrate specificity of the cascade by the covalent attachment of ubiquitin to substrate proteins. Currently, there are over 600 putative E3 ligases, but many are poorly characterized, particularly with respect to individual protein substrates. Here, we highlight systematic approaches to identify and validate UPS targets and discuss how they are underpinning rapid advances in our understanding of the biochemistry and biology of the UPS. The integration of novel tools, model systems and methods for target identification is driving significant interest in drug development, targeting various aspects of UPS function and advancing the understanding of a diverse range of disease processes.
Keywords: cancer; functional genomics; neurodegeneration; proteomics; proteostasis; ubiquitin.
© 2016 The Author(s).
Figures
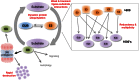
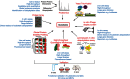
References
-
- Chau V., Tobias J.W., Bachmair A., Marriott D., Ecker D.J., Gonda D.K. et al. (1989) A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243, 1576–1583 PMID: - PubMed
-
- Koepp D.M., Harper J.W. and Elledge S.J. (1999) How the cyclin became a cyclin: regulated proteolysis in the cell cycle. Cell 97, 431–434 doi:http://dx.doi.org/10.1016/S0092-8674(00)80753-9 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

