Heterochromatin assembly by interrupted Sir3 bridges across neighboring nucleosomes
- PMID: 27835568
- PMCID: PMC5106214
- DOI: 10.7554/eLife.17556
Heterochromatin assembly by interrupted Sir3 bridges across neighboring nucleosomes
Abstract
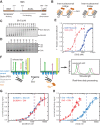
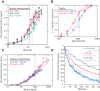
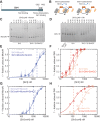
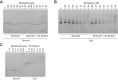
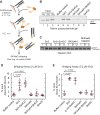
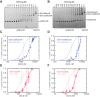
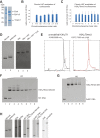
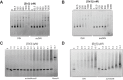
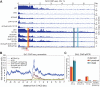
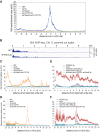
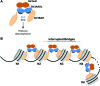
Heterochromatin is a conserved feature of eukaryotic chromosomes with central roles in regulation of gene expression and maintenance of genome stability. Heterochromatin formation involves spreading of chromatin-modifying factors away from initiation points over large DNA domains by poorly understood mechanisms. In Saccharomyces cerevisiae, heterochromatin formation requires the SIR complex, which contains subunits with histone-modifying, histone-binding, and self-association activities. Here, we analyze binding of the Sir proteins to reconstituted mono-, di-, tri-, and tetra-nucleosomal chromatin templates and show that key Sir-Sir interactions bridge only sites on different nucleosomes but not sites on the same nucleosome, and are therefore 'interrupted' with respect to sites on the same nucleosome. We observe maximal binding affinity and cooperativity to unmodified di-nucleosomes and propose that nucleosome pairs bearing unmodified histone H4-lysine16 and H3-lysine79 form the fundamental units of Sir chromatin binding and that cooperative binding requiring two appropriately modified nucleosomes mediates selective Sir recruitment and spreading.
Keywords: S. cerevisiae; Sir3; Sir4; biochemistry; chromatin spreading; chromosomes; genes; heterochromatin; nucleosome.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures





References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

