Identification of SUMO modification sites in the base excision repair protein, Ntg1
- PMID: 27839712
- PMCID: PMC5133457
- DOI: 10.1016/j.dnarep.2016.10.011
Identification of SUMO modification sites in the base excision repair protein, Ntg1
Abstract
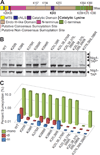
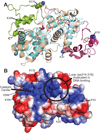
DNA damaging agents are a constant threat to genomes in both the nucleus and the mitochondria. To combat this threat, a suite of DNA repair pathways cooperate to repair numerous types of DNA damage. If left unrepaired, these damages can result in the accumulation of mutations which can lead to deleterious consequences including cancer and neurodegenerative disorders. The base excision repair (BER) pathway is highly conserved from bacteria to humans and is primarily responsible for the removal and subsequent repair of toxic and mutagenic oxidative DNA lesions. Although the biochemical steps that occur in the BER pathway have been well defined, little is known about how the BER machinery is regulated. The budding yeast, Saccharomyces cerevisiae is a powerful model system to biochemically and genetically dissect BER. BER is initiated by DNA N-glycosylases, such as S. cerevisiae Ntg1. Previous work demonstrates that Ntg1 is post-translationally modified by SUMO in response to oxidative DNA damage suggesting that this modification could modulate the function of Ntg1. In this study, we mapped the specific sites of SUMO modification within Ntg1 and identified the enzymes responsible for sumoylating/desumoylating Ntg1. Using a non-sumoylatable version of Ntg1, ntg1ΔSUMO, we performed an initial assessment of the functional impact of Ntg1 SUMO modification in the cellular response to DNA damage. Finally, we demonstrate that, similar to Ntg1, the human homologue of Ntg1, NTHL1, can also be SUMO-modified in response to oxidative stress. Our results suggest that SUMO modification of BER proteins could be a conserved mechanism to coordinate cellular responses to DNA damage.
Keywords: Base excision repair (BER); NTHL1; Ntg1; SUMO; Sumoylation.
Copyright © 2016 Elsevier B.V. All rights reserved.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures



References
-
- Altieri F, Grillo C, Maceroni M, Chichiarelli S. DNA damage and repair: from molecular mechanisms to health implications. Antioxid Redox Signal. 2008;10(5):891–937. - PubMed
-
- Fortini P, Pascucci B, Parlanti E, D'Errico M, Simonelli V, Dogliotti E. The base excision repair: mechanisms and its relevance for cancer susceptibility. Biochimie. 2003;85(11):1053–1071. - PubMed
-
- Boesch P, Weber-Lotfi F, Ibrahim N, Tarasenko V, Cosset A, Paulus F, et al. DNA repair in organelles: Pathways, organization, regulation, relevance in disease and aging. Biochim Biophys Acta. 2011;1813(1):186–200. - PubMed
-
- Kryston TB, Georgiev AB, Pissis P, Georgakilas AG. Role of oxidative stress and DNA damage in human carcinogenesis. Mutation research. 2011;711(1–2):193–201. - PubMed
-
- Pan Y. Mitochondria, reactive oxygen species, and chronological aging: a message from yeast. Experimental gerontology. 2011;46(11):847–852. - PubMed
Web References
-
- http://www.abgent.com/sumoplot Date Last used: 8/4/16.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

