Pre-mRNA Processing Factor Prp18 Is a Stimulatory Factor of Influenza Virus RNA Synthesis and Possesses Nucleoprotein Chaperone Activity
- PMID: 27852861
- PMCID: PMC5244316
- DOI: 10.1128/JVI.01398-16
Pre-mRNA Processing Factor Prp18 Is a Stimulatory Factor of Influenza Virus RNA Synthesis and Possesses Nucleoprotein Chaperone Activity
Abstract
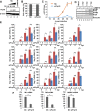
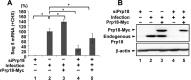
The genome of influenza virus (viral RNA [vRNA]) is associated with the nucleoprotein (NP) and viral RNA-dependent RNA polymerases and forms helical viral ribonucleoprotein (vRNP) complexes. The NP-vRNA complex is the biologically active template for RNA synthesis by the viral polymerase. Previously, we identified human pre-mRNA processing factor 18 (Prp18) as a stimulatory factor for viral RNA synthesis using a Saccharomyces cerevisiae replicon system and a single-gene deletion library of Saccharomyces cerevisiae (T. Naito, Y. Kiyasu, K. Sugiyama, A. Kimura, R. Nakano, A. Matsukage, and K. Nagata, Proc Natl Acad Sci USA, 104:18235-18240, 2007, https://doi.org/10.1073/pnas.0705856104). In infected Prp18 knockdown (KD) cells, the synthesis of vRNA, cRNA, and viral mRNAs was reduced. Prp18 was found to stimulate in vitro viral RNA synthesis through its interaction with NP. Analyses using in vitro RNA synthesis reactions revealed that Prp18 dissociates newly synthesized RNA from the template after the early elongation step to stimulate the elongation reaction. We found that Prp18 functions as a chaperone for NP to facilitate the formation of NP-RNA complexes. Based on these results, it is suggested that Prp18 accelerates influenza virus RNA synthesis as an NP chaperone for the processive elongation reaction.
Importance: Templates for viral RNA synthesis of negative-stranded RNA viruses are not naked RNA but rather RNA encapsidated by viral nucleocapsid proteins forming vRNP complexes. However, viral basic proteins tend to aggregate under physiological ionic strength without chaperones. We identified the pre-mRNA processing factor Prp18 as a stimulatory factor for influenza virus RNA synthesis. We found that one of the targets of Prp18 is NP. Prp18 facilitates the elongation reaction of viral polymerases by preventing the deleterious annealing of newly synthesized RNA to the template. Prp18 functions as a chaperone for NP to stimulate the formation of NP-RNA complexes. Based on these results, we propose that Prp18 may be required to maintain the structural integrity of vRNP for processive template reading.
Keywords: host factor; protein chaperones; ribonucleoprotein; viral RNA synthesis.
Copyright © 2017 American Society for Microbiology.
Figures





Similar articles
-
Cellular splicing factor UAP56 stimulates trimeric NP formation for assembly of functional influenza viral ribonucleoprotein complexes.Sci Rep. 2017 Oct 25;7(1):14053. doi: 10.1038/s41598-017-13784-4. Sci Rep. 2017. PMID: 29070793 Free PMC article.
-
The Host Factor ANP32A Is Required for Influenza A Virus vRNA and cRNA Synthesis.J Virol. 2022 Feb 23;96(4):e0209221. doi: 10.1128/jvi.02092-21. Epub 2021 Dec 22. J Virol. 2022. PMID: 34935435 Free PMC article.
-
Ultrastructure of influenza virus ribonucleoprotein complexes during viral RNA synthesis.Commun Biol. 2021 Jul 9;4(1):858. doi: 10.1038/s42003-021-02388-4. Commun Biol. 2021. PMID: 34244608 Free PMC article.
-
[Dynamics of the influenza virus genome regulated by cellular host factors].Uirusu. 2017;67(1):59-68. doi: 10.2222/jsv.67.59. Uirusu. 2017. PMID: 29593154 Review. Japanese.
-
Emerging roles for the influenza A virus nuclear export protein (NEP).PLoS Pathog. 2012;8(12):e1003019. doi: 10.1371/journal.ppat.1003019. Epub 2012 Dec 6. PLoS Pathog. 2012. PMID: 23236273 Free PMC article. Review.
Cited by
-
Assembly and remodeling of viral DNA and RNA replicons regulated by cellular molecular chaperones.Biophys Rev. 2018 Apr;10(2):445-452. doi: 10.1007/s12551-017-0333-z. Epub 2017 Nov 22. Biophys Rev. 2018. PMID: 29170971 Free PMC article. Review.
-
Differential gene expression reveals host factors for viral shedding variation in mallards (Anas platyrhynchos) infected with low-pathogenic avian influenza virus.J Gen Virol. 2022 Mar;103(3):10.1099/jgv.0.001724. doi: 10.1099/jgv.0.001724. J Gen Virol. 2022. PMID: 35353676 Free PMC article.
-
Fundamental Contribution and Host Range Determination of ANP32A and ANP32B in Influenza A Virus Polymerase Activity.J Virol. 2019 Jun 14;93(13):e00174-19. doi: 10.1128/JVI.00174-19. Print 2019 Jul 1. J Virol. 2019. PMID: 30996088 Free PMC article.
-
Cellular splicing factor UAP56 stimulates trimeric NP formation for assembly of functional influenza viral ribonucleoprotein complexes.Sci Rep. 2017 Oct 25;7(1):14053. doi: 10.1038/s41598-017-13784-4. Sci Rep. 2017. PMID: 29070793 Free PMC article.
-
Transcriptome profiling of human oocytes experiencing recurrent total fertilization failure.Sci Rep. 2018 Dec 17;8(1):17890. doi: 10.1038/s41598-018-36275-6. Sci Rep. 2018. PMID: 30559372 Free PMC article.
References
-
- Yamanaka K, Ishihama A, Nagata K. 1990. Reconstitution of influenza virus RNA-nucleoprotein complexes structurally resembling native viral ribonucleoprotein cores. J Biol Chem 265:11151–11155. - PubMed
-
- Honda A, Ueda K, Nagata K, Ishihama A. 1988. RNA polymerase of influenza virus: role of NP in RNA chain elongation. J Biochem 104:1021–1026. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

