Local Arp2/3-dependent actin assembly modulates applied traction force during apCAM adhesion site maturation
- PMID: 27852899
- PMCID: PMC5221634
- DOI: 10.1091/mbc.E16-04-0228
Local Arp2/3-dependent actin assembly modulates applied traction force during apCAM adhesion site maturation
Abstract
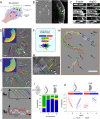
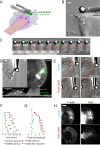
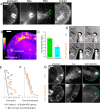
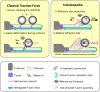
Homophilic binding of immunoglobulin superfamily molecules such as the Aplysia cell adhesion molecule (apCAM) leads to actin filament assembly near nascent adhesion sites. Such actin assembly can generate significant localized forces that have not been characterized in the larger context of axon growth and guidance. We used apCAM-coated bead substrates applied to the surface of neuronal growth cones to characterize the development of forces evoked by varying stiffness of mechanical restraint. Unrestrained bead propulsion matched or exceeded rates of retrograde network flow and was dependent on Arp2/3 complex activity. Analysis of growth cone forces applied to beads at low stiffness of restraint revealed switching between two states: frictional coupling to retrograde flow and Arp2/3-dependent propulsion. Stiff mechanical restraint led to formation of an extensive actin cup matching the geometric profile of the bead target and forward growth cone translocation; pharmacological inhibition of the Arp2/3 complex or Rac attenuated F-actin assembly near bead binding sites, decreased the efficacy of growth responses, and blocked accumulation of signaling molecules associated with nascent adhesions. These studies introduce a new model for regulation of traction force in which local actin assembly forces buffer nascent adhesion sites from the mechanical effects of retrograde flow.
© 2017 Buck et al. This article is distributed by The American Society for Cell Biology under license from the author(s). Two months after publication it is available to the public under an Attribution–Noncommercial–Share Alike 3.0 Unported Creative Commons License (http://creativecommons.org/licenses/by-nc-sa/3.0).
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

