Substrate recognition and catalysis by GH47 α-mannosidases involved in Asn-linked glycan maturation in the mammalian secretory pathway
- PMID: 27856750
- PMCID: PMC5150396
- DOI: 10.1073/pnas.1611213113
Substrate recognition and catalysis by GH47 α-mannosidases involved in Asn-linked glycan maturation in the mammalian secretory pathway
Abstract
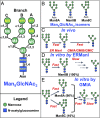
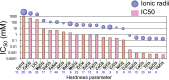
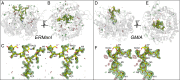
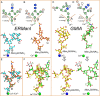
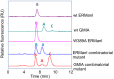
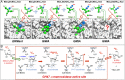
Maturation of Asn-linked oligosaccharides in the eukaryotic secretory pathway requires the trimming of nascent glycan chains to remove all glucose and several mannose residues before extension into complex-type structures on the cell surface and secreted glycoproteins. Multiple glycoside hydrolase family 47 (GH47) α-mannosidases, including endoplasmic reticulum (ER) α-mannosidase I (ERManI) and Golgi α-mannosidase IA (GMIA), are responsible for cleavage of terminal α1,2-linked mannose residues to produce uniquely trimmed oligomannose isomers that are necessary for ER glycoprotein quality control and glycan maturation. ERManI and GMIA have similar catalytic domain structures, but each enzyme cleaves distinct residues from tribranched oligomannose glycan substrates. The structural basis for branch-specific cleavage by ERManI and GMIA was explored by replacing an essential enzyme-bound Ca2+ ion with a lanthanum (La3+) ion. This ion swap led to enzyme inactivation while retaining high-affinity substrate interactions. Cocrystallization of La3+-bound enzymes with Man9GlcNAc2 substrate analogs revealed enzyme-substrate complexes with distinct modes of glycan branch insertion into the respective enzyme active-site clefts. Both enzymes had glycan interactions that extended across the entire glycan structure, but each enzyme engaged a different glycan branch and used different sets of glycan interactions. Additional mutagenesis and time-course studies of glycan cleavage probed the structural basis of enzyme specificity. The results provide insights into the enzyme catalytic mechanisms and reveal structural snapshots of the sequential glycan cleavage events. The data also indicate that full steric access to glycan substrates determines the efficiency of mannose-trimming reactions that control the conversion to complex-type structures in mammalian cells.
Keywords: Asn-linked glycan processing; bioinorganic chemistry; glycosidase mechanism; glycoside hydrolase; α-mannosidase.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Family 47 alpha-mannosidases in N-glycan processing.Methods Enzymol. 2006;415:31-46. doi: 10.1016/S0076-6879(06)15003-X. Methods Enzymol. 2006. PMID: 17116466 Free PMC article.
-
Identification, expression, and characterization of a cDNA encoding human endoplasmic reticulum mannosidase I, the enzyme that catalyzes the first mannose trimming step in mammalian Asn-linked oligosaccharide biosynthesis.J Biol Chem. 1999 Jul 23;274(30):21375-86. doi: 10.1074/jbc.274.30.21375. J Biol Chem. 1999. PMID: 10409699
-
Characterization of a cDNA encoding a novel human Golgi alpha 1, 2-mannosidase (IC) involved in N-glycan biosynthesis.J Biol Chem. 2000 Oct 13;275(41):31655-60. doi: 10.1074/jbc.M004935200. J Biol Chem. 2000. PMID: 10915796
-
Importance of glycosidases in mammalian glycoprotein biosynthesis.Biochim Biophys Acta. 1999 Dec 6;1473(1):96-107. doi: 10.1016/s0304-4165(99)00171-3. Biochim Biophys Acta. 1999. PMID: 10580131 Review.
-
Golgi alpha-mannosidase II deficiency in vertebrate systems: implications for asparagine-linked oligosaccharide processing in mammals.Biochim Biophys Acta. 2002 Dec 19;1573(3):225-35. doi: 10.1016/s0304-4165(02)00388-4. Biochim Biophys Acta. 2002. PMID: 12417404 Review.
Cited by
-
Exploitation of glycosylation in enveloped virus pathobiology.Biochim Biophys Acta Gen Subj. 2019 Oct;1863(10):1480-1497. doi: 10.1016/j.bbagen.2019.05.012. Epub 2019 May 20. Biochim Biophys Acta Gen Subj. 2019. PMID: 31121217 Free PMC article. Review.
-
Elevated high-mannose N-glycans hamper endometrial decidualization.iScience. 2023 Oct 14;26(11):108170. doi: 10.1016/j.isci.2023.108170. eCollection 2023 Nov 17. iScience. 2023. PMID: 37915610 Free PMC article.
-
Resveratrol targets PD-L1 glycosylation and dimerization to enhance antitumor T-cell immunity.Aging (Albany NY). 2020 Jan 4;12(1):8-34. doi: 10.18632/aging.102646. Epub 2020 Jan 4. Aging (Albany NY). 2020. PMID: 31901900 Free PMC article.
-
Perspective on Immunoglobulin N-Glycosylation Status in Follicular Lymphoma: Uncovering BCR-Dependent and Independent Mechanisms Driving Subclonal Evolution.Cancers (Basel). 2025 Apr 4;17(7):1219. doi: 10.3390/cancers17071219. Cancers (Basel). 2025. PMID: 40227774 Free PMC article.
-
Modulation of ERQC and ERAD: A Broad-Spectrum Spanner in the Works of Cancer Cells?J Oncol. 2019 Oct 1;2019:8384913. doi: 10.1155/2019/8384913. eCollection 2019. J Oncol. 2019. PMID: 31662755 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

