Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity
- PMID: 27857054
- PMCID: PMC5143421
- DOI: 10.1038/cr.2016.135
Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity
Abstract
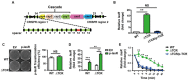
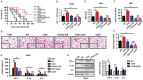
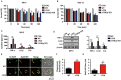
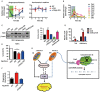
Clustered regularly interspaced short palindromic repeats (CRISPR)-CRISPR-associated (Cas) systems in bacteria and archaea provide adaptive immunity against invading foreign nucleic acids. Previous studies suggest that certain bacteria employ their Type II CRISPR-Cas systems to target their own genes, thus evading host immunity. However, whether other CRISPR-Cas systems have similar functions during bacterial invasion of host cells remains unknown. Here we identify a novel role for Type I CRISPR-Cas systems in evading host defenses in Pseudomonas aeruginosa strain UCBPP-PA14. The Type I CRISPR-Cas system of PA14 targets the mRNA of the bacterial quorum-sensing regulator LasR to dampen the recognition by toll-like receptor 4, thus diminishing the pro-inflammatory responses of the host in cell and mouse models. Mechanistically, this nuclease-mediated RNA degradation requires a "5'-GGN-3'" recognition motif in the target mRNA, and HD and DExD/H domains in Cas3 of the Type I CRISPR-Cas system. As LasR and Type I CRISPR-Cas systems are ubiquitously present in bacteria, our findings elucidate an important common mechanism underlying bacterial virulence.
Figures


Comment in
-
CRISPR control of virulence in Pseudomonas aeruginosa.Cell Res. 2017 Feb;27(2):163-164. doi: 10.1038/cr.2017.6. Epub 2017 Jan 13. Cell Res. 2017. PMID: 28084330 Free PMC article.
-
Commentary: Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity.Front Microbiol. 2017 Feb 28;8:319. doi: 10.3389/fmicb.2017.00319. eCollection 2017. Front Microbiol. 2017. PMID: 28293232 Free PMC article. No abstract available.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

