Tanshinone IIA combined with adriamycin inhibited malignant biological behaviors of NSCLC A549 cell line in a synergistic way
- PMID: 27863471
- PMCID: PMC5116215
- DOI: 10.1186/s12885-016-2921-x
Tanshinone IIA combined with adriamycin inhibited malignant biological behaviors of NSCLC A549 cell line in a synergistic way
Abstract
Background: The study was designed to develop a platform to verify whether the extract of herbs combined with chemotherapy drugs play a synergistic role in anti-tumor effects, and to provide experimental evidence and theoretical reference for finding new effective sensitizers.
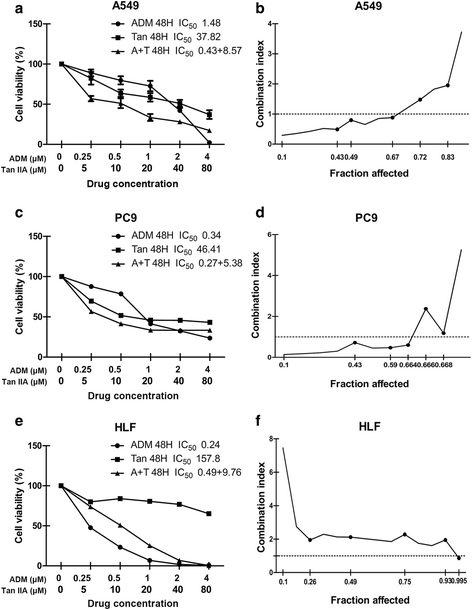
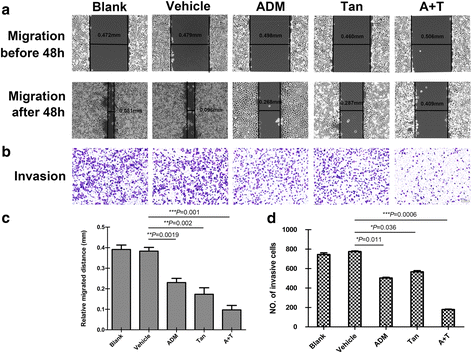
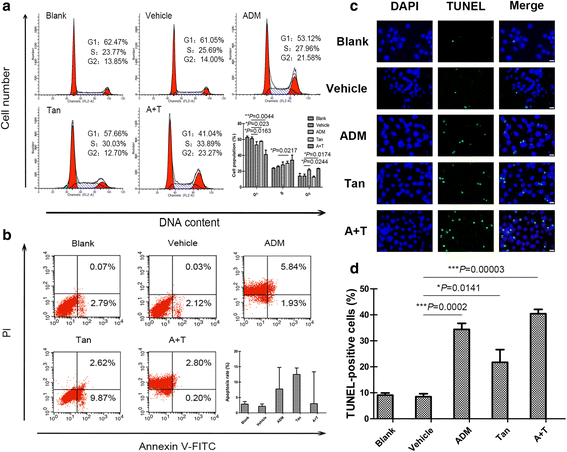
Methods: Inhibition of tanshinone IIA and adriamycin on the proliferation of A549, PC9 and HLF cells were assessed by CCK8 assays. The combination index (CI) was calculated with the Chou-Talalay method, based on the median-effect principle. Migration and invasion ability of A549 cells were determined by wound healing assay and transwell assay. Flow cytometry was used to detect the cell apoptosis and the distribution of cell cycles. TUNEL staining was used to detect the apoptotic cells. Immunofluorescence staining was used to detect the expression of Cleaved Caspase-3. Western blotting was used to detect the proteins expression of relative apoptotic signal pathways. CDOCKER module in DS 2.5 was used to detect the binding modes of the drugs and the proteins.
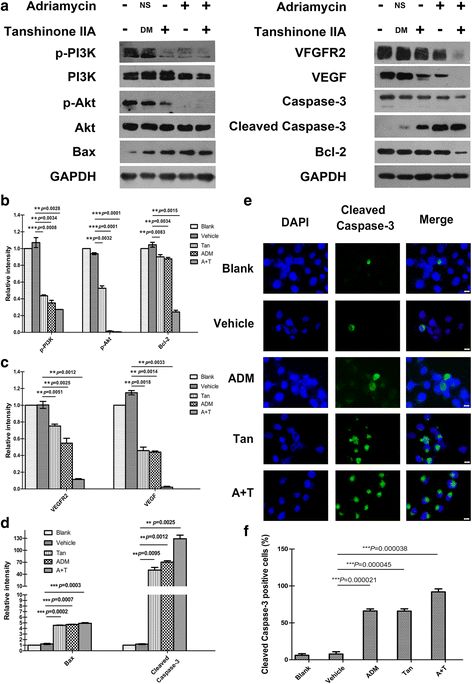
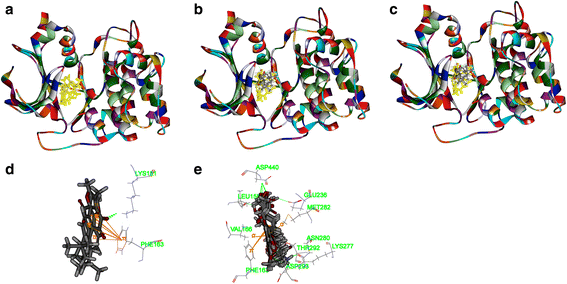
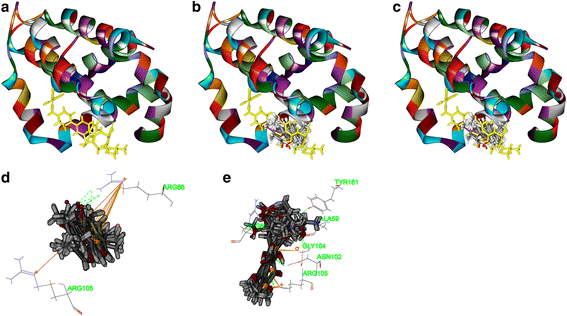
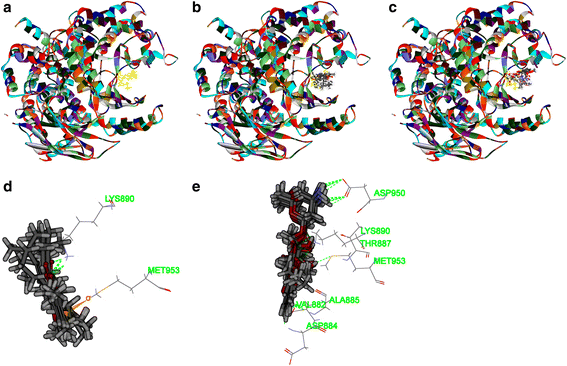
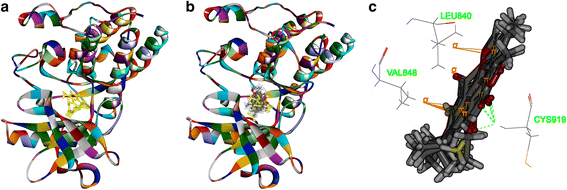
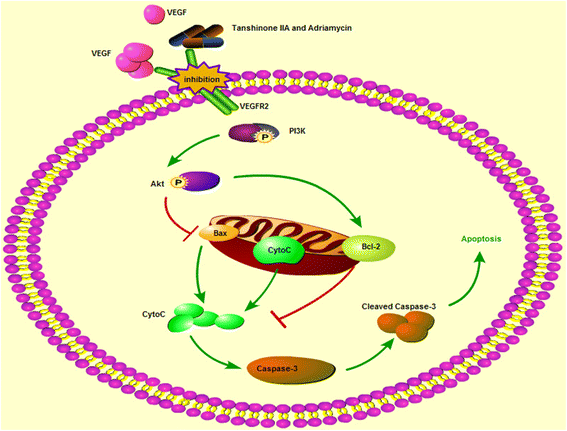
Results: Both tanshinone IIA and adriamycin could inhibit the growth of A549, PC9, and HLF cells in a dose- and time-dependent manner, while the proliferative inhibition effect of tanshinone IIA on cells was much weaker than that of adriamycin. Different from the cancer cells, HLF cells displayed a stronger sensitivity to adriamycin, and a weaker sensitivity to tanshinone IIA. When tanshinone IIA combined with adriamycin at a ratio of 20:1, they exhibited a synergistic anti-proliferation effect on A549 and PC9 cells, but not in HLF cells. Tanshinone IIA combined with adriamycin could synergistically inhibit migration, induce apoptosis and arrest cell cycle at the S and G2 phases in A549 cells. Both groups of the single drug treatment and the drug combination up-regulated the expressions of Cleaved Caspase-3 and Bax, but down-regulated the expressions of VEGF, VEGFR2, p-PI3K, p-Akt, Bcl-2, and Caspase-3 protein. Compared with the single drug treatment groups, the drug combination groups were more statistically significant. The molecular docking algorithms indicated that tanshinone IIA could be docked into the active sites of all the tested proteins with H-bond and aromatic interactions, compared with that of adriamycin.
Conclusions: Tanshinone IIA can be developed as a novel agent in the postoperative adjuvant therapy combined with other anti-tumor agents, and improve the sensibility of chemotherapeutics for non-small cell lung cancer with fewer side effects. In addition, this experiment can not only provide a reference for the development of more effective anti-tumor medicine ingredients, but also build a platform for evaluating the anti-tumor effects of Chinese herbal medicines in combination with chemotherapy drugs.
Keywords: A549; Adriamycin; NSCLC; Synergistic effect; Tanshinone IIA; VEGF/PI3K/Akt signal pathway.
Figures
References
-
- Nagel R, Stigter-van Walsum M, Buijze M, van den Berg J, van der Meulen IH, Hodzic J, Piersma SR, Pham TV, Jimenez CR, van Beusechem VW, et al. Genome-wide siRNA Screen Identifies the Radiosensitizing Effect of Downregulation of MASTL and FOXM1 in NSCLC. Mol Cancer Ther. 2015;14(6):1434–1444. doi: 10.1158/1535-7163.MCT-14-0846. - DOI - PubMed
-
- Bunn PA, Jr, Thatcher N. Systemic treatment for advanced (stage IIIb/IV) non-small cell lung cancer: more treatment options; more things to consider. Conclusion. Oncologist. 2008;37(Suppl 1):46. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

