Post-translational Control of Intracellular Pathogen Sensing Pathways
- PMID: 27863906
- PMCID: PMC5580928
- DOI: 10.1016/j.it.2016.10.008
Post-translational Control of Intracellular Pathogen Sensing Pathways
Abstract
Mammalian cells recognize virus-derived nucleic acids using a defined set of intracellular sensors including the DNA sensors cyclic GMP-AMP (cGAMP) synthase (cGAS) and interferon gamma (IFNγ)-inducible protein 16 (IFI16) as well as viral RNA receptors of the retinoic acid-inducible gene I (RIG-I)-like receptor (RLR) family. Following innate immune recognition, these sensors launch an immune response that is characterized by the transcriptional upregulation of many antiviral molecules, including proinflammatory cytokines, chemokines, and IFN-stimulated genes. Recent studies have demonstrated that the signal transduction initiated by these sensors is sophisticatedly regulated by post-translational modifications (PTMs) resulting in a robust yet 'tunable' cytokine response to maintain immune homeostasis. Here we summarize recent advances in our understanding of how PTMs and regulatory enzymes control the signaling activity of RLRs, cGAS, and IFI16 as well as their proximal adaptor proteins.
Copyright © 2016 Elsevier Ltd. All rights reserved.
Figures
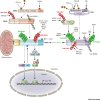

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

