Exploring the Role of the Ω-Loop in the Evolution of Ceftazidime Resistance in the PenA β-Lactamase from Burkholderia multivorans, an Important Cystic Fibrosis Pathogen
- PMID: 27872073
- PMCID: PMC5278747
- DOI: 10.1128/AAC.01941-16
Exploring the Role of the Ω-Loop in the Evolution of Ceftazidime Resistance in the PenA β-Lactamase from Burkholderia multivorans, an Important Cystic Fibrosis Pathogen
Abstract
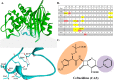
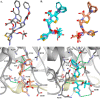
The unwelcome evolution of resistance to the advanced generation cephalosporin antibiotic, ceftazidime is hindering the effective therapy of Burkholderia cepacia complex (BCC) infections. Regrettably, BCC organisms are highly resistant to most antibiotics, including polymyxins; ceftazidime and trimethoprim-sulfamethoxazole are the most effective treatment options. Unfortunately, resistance to ceftazidime is increasing and posing a health threat to populations susceptible to BCC infection. We found that up to 36% of 146 tested BCC clinical isolates were nonsusceptible to ceftazidime (MICs ≥ 8 μg/ml). To date, the biochemical basis for ceftazidime resistance in BCC is largely undefined. In this study, we investigated the role of the Ω-loop in mediating ceftazidime resistance in the PenA β-lactamase from Burkholderia multivorans, a species within the BCC. Single amino acid substitutions were engineered at selected positions (R164, T167, L169, and D179) in the PenA β-lactamase. Cell-based susceptibility testing revealed that 21 of 75 PenA variants engineered in this study were resistant to ceftazidime, with MICs of >8 μg/ml. Under steady-state conditions, each of the selected variants (R164S, T167G, L169A, and D179N) demonstrated a substrate preference for ceftazidime compared to wild-type PenA (32- to 320-fold difference). Notably, the L169A variant hydrolyzed ceftazidime significantly faster than PenA and possessed an ∼65-fold-lower apparent Ki (Kiapp) than that of PenA. To understand why these amino acid substitutions result in enhanced ceftazidime binding and/or turnover, we employed molecular dynamics simulation (MDS). The MDS suggested that the L169A variant starts with the most energetically favorable conformation (-28.1 kcal/mol), whereas PenA possessed the most unfavorable initial conformation (136.07 kcal/mol). In addition, we observed that the spatial arrangement of E166, N170, and the hydrolytic water molecules may be critical for enhanced ceftazidime hydrolysis by the L169A variant. Importantly, we found that two clinical isolates of B. multivorans possessed L169 amino acid substitutions (L169F and L169P) in PenA and were highly resistant to ceftazidime (MICs ≥ 512 μg/ml). In conclusion, substitutions in the Ω-loop alter the positioning of the hydrolytic machinery as well as allow for a larger opening of the active site to accommodate the bulky R1 and R2 side chains of ceftazidime, resulting in resistance. This analysis provides insights into the emerging phenotype of ceftazidime-resistant BCC and explains the evolution of amino acid substitutions in the Ω-loop of PenA of this significant clinical pathogen.
Keywords: Burkholderia; beta-lactamase; beta-lactams; ceftazidime.
Copyright © 2017 American Society for Microbiology.
Figures



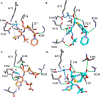
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

