Predicting the pathogenicity of aminoacyl-tRNA synthetase mutations
- PMID: 27876679
- PMCID: PMC5253330
- DOI: 10.1016/j.ymeth.2016.11.013
Predicting the pathogenicity of aminoacyl-tRNA synthetase mutations
Abstract
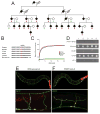
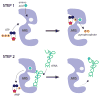
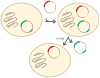
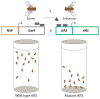
Aminoacyl-tRNA synthetases (ARSs) are ubiquitously expressed, essential enzymes responsible for charging tRNA with cognate amino acids-the first step in protein synthesis. ARSs are required for protein translation in the cytoplasm and mitochondria of all cells. Surprisingly, mutations in 28 of the 37 nuclear-encoded human ARS genes have been linked to a variety of recessive and dominant tissue-specific disorders. Current data indicate that impaired enzyme function is a robust predictor of the pathogenicity of ARS mutations. However, experimental model systems that distinguish between pathogenic and non-pathogenic ARS variants are required for implicating newly identified ARS mutations in disease. Here, we outline strategies to assist in predicting the pathogenicity of ARS variants and urge cautious evaluation of genetic and functional data prior to linking an ARS mutation to a human disease phenotype.
Keywords: Aminoacyl-tRNA synthetases; Charcot-Marie-Tooth (CMT) disease; Functional evaluation of disease-associated mutations; Mendelian disease; Neurodevelopmental disease; Peripheral neuropathy.
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

