Multi-omics Analysis of Serum Samples Demonstrates Reprogramming of Organ Functions Via Systemic Calcium Mobilization and Platelet Activation in Metastatic Melanoma
- PMID: 27879288
- PMCID: PMC5217785
- DOI: 10.1074/mcp.M116.063313
Multi-omics Analysis of Serum Samples Demonstrates Reprogramming of Organ Functions Via Systemic Calcium Mobilization and Platelet Activation in Metastatic Melanoma
Abstract
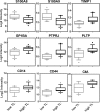
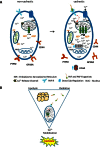
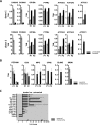
Pathophysiologies of cancer-associated syndromes such as cachexia are poorly understood and no routine biomarkers have been established, yet. Using shotgun proteomics, known marker molecules including PMEL, CRP, SAA, and CSPG4 were found deregulated in patients with metastatic melanoma. Targeted analysis of 58 selected proteins with multiple reaction monitoring was applied for independent data verification. In three patients, two of which suffered from cachexia, a tissue damage signature was determined, consisting of nine proteins, PLTP, CD14, TIMP1, S10A8, S10A9, GP1BA, PTPRJ, CD44, and C4A, as well as increased levels of glycine and asparagine, and decreased levels of polyunsaturated phosphatidylcholine concentrations, as determined by targeted metabolomics. Remarkably, these molecules are known to be involved in key processes of cancer cachexia. Based on these results, we propose a model how metastatic melanoma may lead to reprogramming of organ functions via formation of platelet activating factors from long-chain polyunsaturated phosphatidylcholines under oxidative conditions and via systemic induction of intracellular calcium mobilization. Calcium mobilization in platelets was demonstrated to alter levels of several of these marker molecules. Additionally, platelets from melanoma patients proved to be in a rather exhausted state, and platelet-derived eicosanoids implicated in tumor growth were found massively increased in blood from three melanoma patients. Platelets were thus identified as important source of serum protein and lipid alterations in late stage melanoma patients. As a result, the proposed model describes the crosstalk between lipolysis of fat tissue and muscle wasting mediated by oxidative stress, resulting in the metabolic deregulations characteristic for cachexia.
© 2017 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


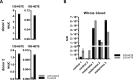
References
-
- Adkins J. N., Varnum S. M., Auberry K. J., Moore R. J., Angell N. H., Smith R. D., Springer D. L., and Pounds J. G. (2002) Toward a human blood serum proteome: analysis by multidimensional separation coupled with mass spectrometry. Mol. Cell. Proteomics 1, 947–955 - PubMed
-
- Abbatiello S. E., Schilling B., Mani D. R., Zimmerman L. J., Hall S. C., MacLean B., Albertolle M., Allen S., Burgess M., Cusack M. P., Gosh M., Hedrick V., Held J. M., Inerowicz H. D., Jackson A., Keshishian H., Kinsinger C. R., Lyssand J., Makowski L., Mesri M., Rodriguez H., Rudnick P., Sadowski P., Sedransk N., Shaddox K., Skates S. J., Kuhn E., Smith D., Whiteaker J. R., Whitwell C., Zhang S., Borchers C. H., Fisher S. J., Gibson B. W., Liebler D. C., MacCoss M. J., Neubert T. A., Paulovich A. G., Regnier F. E., Tempst P., and Carr S. A. (2015) Large-Scale Interlaboratory Study to Develop, Analytically Validate and Apply Highly Multiplexed, Quantitative Peptide Assays to Measure Cancer-Relevant Proteins in Plasma. Mol. Cell. Proteomics 14, 2357–2374 - PMC - PubMed
-
- Paulitschke V., Berger W., Paulitschke P., Hofstatter E., Knapp B., Dingelmaier-Hovorka R., Fodinger D., Jager W., Szekeres T., Meshcheryakova A., Bileck A., Pirker C., Pehamberger H., Gerner C., and Kunstfeld R. (2015) Vemurafenib resistance signature by proteome analysis offers new strategies and rational therapeutic concepts. Mol. Cancer Therap. 14, 757–768 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

