CENP-A Is Dispensable for Mitotic Centromere Function after Initial Centromere/Kinetochore Assembly
- PMID: 27880912
- PMCID: PMC5134894
- DOI: 10.1016/j.celrep.2016.10.084
CENP-A Is Dispensable for Mitotic Centromere Function after Initial Centromere/Kinetochore Assembly
Abstract
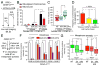
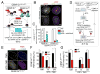
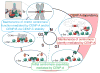
Human centromeres are defined by chromatin containing the histone H3 variant CENP-A assembled onto repetitive alphoid DNA sequences. By inducing rapid, complete degradation of endogenous CENP-A, we now demonstrate that once the first steps of centromere assembly have been completed in G1/S, continued CENP-A binding is not required for maintaining kinetochore attachment to centromeres or for centromere function in the next mitosis. Degradation of CENP-A prior to kinetochore assembly is found to block deposition of CENP-C and CENP-N, but not CENP-T, thereby producing defective kinetochores and failure of chromosome segregation. Without the continuing presence of CENP-A, CENP-B binding to alphoid DNA sequences becomes essential to preserve anchoring of CENP-C and the kinetochore to each centromere. Thus, there is a reciprocal interdependency of CENP-A chromatin and the underlying repetitive centromere DNA sequences bound by CENP-B in the maintenance of human chromosome segregation.
Keywords: CENP-A; CENP-B; CENP-C; auxin; centromere; chromosome segregation; epigenetic; kinetochore; mitosis; protein degradation.
Copyright © 2016 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures


References
-
- . CCAN makes multiple contacts with centromeric DNA to provide distinct pathways to the outer kinetochore. 2008;135(6):1039–1052. - PubMed
-
- Bade D, et al. The E3 Ligase CUL3/RDX Controls Centromere Maintenance by Ubiquitylating and Stabilizing CENP-A in a CAL1-Dependent Manner. Developmental Cell. 2014;28(5):508–519. - PubMed
-
- Black BE, et al. Centromere Identity Maintained by Nucleosomes Assembled with Histone H3 Containing the CENP-A Targeting Domain. Molecular Cell. 2007;25(2):309–322. - PubMed
-
- Black BE, et al. Structural determinants for generating centromeric chromatin. Nature. 2004;430(6999):578–582. - PubMed
-
- Bodor DL, et al. Analysis of protein turnover by quantitative SNAP-based pulse-chase imaging. In: Bonifacino Juan S, et al., editors. Current protocols in cell biology / editorial board. Unit8.8 Chapter 8. 2012. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

