Genomic approaches to the assessment of human spina bifida risk
- PMID: 27883265
- PMCID: PMC5388593
- DOI: 10.1002/bdra.23592
Genomic approaches to the assessment of human spina bifida risk
Abstract
Structural birth defects are a leading cause of mortality and morbidity in children world-wide, affecting as much as 6% of all live births. Among these conditions, neural tube defects (NTDs), including spina bifida and anencephaly, arise from a combination of complex gene and environment interactions that are as yet poorly understood within human populations. Rapid advances in massively parallel DNA sequencing and bioinformatics allow for analyses of the entire genome beyond the 2% of the genomic sequence covering protein coding regions. Efforts to collect and analyze these large datasets hold promise for illuminating gene network variations and eventually epigenetic events that increase individual risk for failure to close the neural tube. In this review, we discuss current challenges for DNA genome sequence analysis of NTD affected populations, and compare experience in the field with other complex genetic disorders for which large datasets are accumulating. The ultimate goal of this research is to find strategies for optimizing conditions that promote healthy birth outcomes for individual couples. Birth Defects Research 109:120-128, 2017. © 2016 Wiley Periodicals, Inc.
Keywords: biogeography; complex genetic disorders; intergenic (noncoding) sequence analysis; variant analysis; whole exome sequencing (WES); whole genome sequencing (WGS).
© 2016 Wiley Periodicals, Inc.
Figures
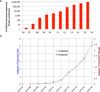
References
-
- Albert FW, Kruglyak L. The role of regulatory variation in complex traits and disease. Nature reviews Genetics. 2015;16:197–212. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

