Satb2 determines miRNA expression and long-term memory in the adult central nervous system
- PMID: 27897969
- PMCID: PMC5207769
- DOI: 10.7554/eLife.17361
Satb2 determines miRNA expression and long-term memory in the adult central nervous system
Abstract
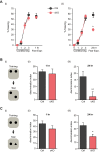
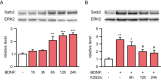
SATB2 is a risk locus for schizophrenia and encodes a DNA-binding protein that regulates higher-order chromatin configuration. In the adult brain Satb2 is almost exclusively expressed in pyramidal neurons of two brain regions important for memory formation, the cerebral cortex and the CA1-hippocampal field. Here we show that Satb2 is required for key hippocampal functions since deletion of Satb2 from the adult mouse forebrain prevents the stabilization of synaptic long-term potentiation and markedly impairs long-term fear and object discrimination memory. At the molecular level, we find that synaptic activity and BDNF up-regulate Satb2, which itself binds to the promoters of coding and non-coding genes. Satb2 controls the hippocampal levels of a large cohort of miRNAs, many of which are implicated in synaptic plasticity and memory formation. Together, our findings demonstrate that Satb2 is critically involved in long-term plasticity processes in the adult forebrain that underlie the consolidation and stabilization of context-linked memory.
Keywords: LTP; chromatin; chromosomes; genes; long-term memory; miRNA; mouse; neuroscience; synaptic plasticity.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures


















References
-
- Balamotis MA, Tamberg N, Woo YJ, Li J, Davy B, Kohwi-Shigematsu T, Kohwi Y. Satb1 ablation alters temporal expression of immediate early genes and reduces dendritic spine density during postnatal brain development. Molecular and Cellular Biology. 2012;32:333–347. doi: 10.1128/MCB.05917-11. - DOI - PMC - PubMed
-
- Baranek C, Dittrich M, Parthasarathy S, Bonnon CG, Britanova O, Lanshakov D, Boukhtouche F, Sommer JE, Colmenares C, Tarabykin V, Atanasoski S. Protooncogene Ski cooperates with the chromatin-remodeling factor Satb2 in specifying callosal neurons. PNAS. 2012;109:3546–3551. doi: 10.1073/pnas.1108718109. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

