Division of labor among Mycobacterium smegmatis RNase H enzymes: RNase H1 activity of RnhA or RnhC is essential for growth whereas RnhB and RnhA guard against killing by hydrogen peroxide in stationary phase
- PMID: 27899559
- PMCID: PMC5224475
- DOI: 10.1093/nar/gkw1046
Division of labor among Mycobacterium smegmatis RNase H enzymes: RNase H1 activity of RnhA or RnhC is essential for growth whereas RnhB and RnhA guard against killing by hydrogen peroxide in stationary phase
Abstract
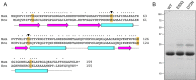
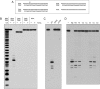
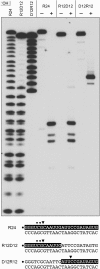
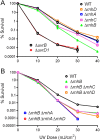
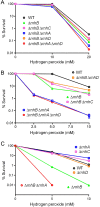
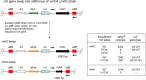
RNase H enzymes sense the presence of ribonucleotides in the genome and initiate their removal by incising the ribonucleotide-containing strand of an RNA:DNA hybrid. Mycobacterium smegmatis encodes four RNase H enzymes: RnhA, RnhB, RnhC and RnhD. Here, we interrogate the biochemical activity and nucleic acid substrate specificity of RnhA. We report that RnhA (like RnhC characterized previously) is an RNase H1-type magnesium-dependent endonuclease with stringent specificity for RNA:DNA hybrid duplexes. Whereas RnhA does not incise an embedded mono-ribonucleotide, it can efficiently cleave within tracts of four or more ribonucleotides in duplex DNA. We gained genetic insights to the division of labor among mycobacterial RNases H by deleting the rnhA, rnhB, rnhC and rnhD genes, individually and in various combinations. The salient conclusions are that: (i) RNase H1 activity is essential for mycobacterial growth and can be provided by either RnhC or RnhA; (ii) the RNase H2 enzymes RnhB and RnhD are dispensable for growth and (iii) RnhB and RnhA collaborate to protect M. smegmatis against oxidative damage in stationary phase. Our findings highlight RnhC, the sole RNase H1 in pathogenic mycobacteria, as a candidate drug discovery target for tuberculosis and leprosy.
© The Author(s) 2016. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
The C-Terminal Acid Phosphatase Module of the RNase HI Enzyme RnhC Controls Rifampin Sensitivity and Light-Dependent Colony Pigmentation of Mycobacterium smegmatis.J Bacteriol. 2023 Apr 25;205(4):e0043122. doi: 10.1128/jb.00431-22. Epub 2023 Mar 14. J Bacteriol. 2023. PMID: 36916909 Free PMC article.
-
Biochemical Characterization of Mycobacterium smegmatis RnhC (MSMEG_4305), a Bifunctional Enzyme Composed of Autonomous N-Terminal Type I RNase H and C-Terminal Acid Phosphatase Domains.J Bacteriol. 2015 Aug 1;197(15):2489-98. doi: 10.1128/JB.00268-15. Epub 2015 May 18. J Bacteriol. 2015. PMID: 25986906 Free PMC article.
-
The deletion of rnhB in Mycobacterium smegmatis does not affect the level of RNase HII substrates or influence genome stability.PLoS One. 2015 Jan 20;10(1):e0115521. doi: 10.1371/journal.pone.0115521. eCollection 2015. PLoS One. 2015. PMID: 25603150 Free PMC article.
-
Synthetic lethal mutants in Escherichia coli define pathways necessary for survival with RNase H deficiency.J Bacteriol. 2023 Oct 26;205(10):e0028023. doi: 10.1128/jb.00280-23. Epub 2023 Oct 11. J Bacteriol. 2023. PMID: 37819120 Free PMC article.
-
RNases H: Structure and mechanism.DNA Repair (Amst). 2019 Dec;84:102672. doi: 10.1016/j.dnarep.2019.102672. Epub 2019 Jul 20. DNA Repair (Amst). 2019. PMID: 31371183 Review.
Cited by
-
The catalytic mechanism, metal dependence, substrate specificity, and biodiversity of ribonuclease H.Front Microbiol. 2022 Nov 21;13:1034811. doi: 10.3389/fmicb.2022.1034811. eCollection 2022. Front Microbiol. 2022. PMID: 36478866 Free PMC article. Review.
-
Functional Disassociation Between the Protein Domains of MSMEG_4305 of Mycolicibacterium smegmatis (Mycobacterium smegmatis) in vivo.Front Microbiol. 2020 Aug 19;11:2008. doi: 10.3389/fmicb.2020.02008. eCollection 2020. Front Microbiol. 2020. PMID: 32973726 Free PMC article.
-
RNase HI Depletion Strongly Potentiates Cell Killing by Rifampicin in Mycobacteria.Antimicrob Agents Chemother. 2022 Oct 18;66(10):e0209121. doi: 10.1128/aac.02091-21. Epub 2022 Sep 26. Antimicrob Agents Chemother. 2022. PMID: 36154174 Free PMC article.
-
The C-Terminal Acid Phosphatase Module of the RNase HI Enzyme RnhC Controls Rifampin Sensitivity and Light-Dependent Colony Pigmentation of Mycobacterium smegmatis.J Bacteriol. 2023 Apr 25;205(4):e0043122. doi: 10.1128/jb.00431-22. Epub 2023 Mar 14. J Bacteriol. 2023. PMID: 36916909 Free PMC article.
-
Comparative genomic analysis identifies potential adaptive variation in Mycoplasma ovipneumoniae.Microb Genom. 2024 Aug;10(8):001279. doi: 10.1099/mgen.0.001279. Microb Genom. 2024. PMID: 39213169 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

